Transcription Factors in Cancer: When Alternative Splicing Determines Opposite Cell Fates
- PMID: 32244895
- PMCID: PMC7140685
- DOI: 10.3390/cells9030760
Transcription Factors in Cancer: When Alternative Splicing Determines Opposite Cell Fates
Abstract
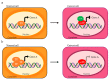
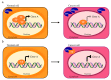
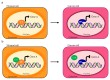
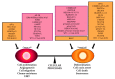
Alternative splicing (AS) is a finely regulated mechanism for transcriptome and proteome diversification in eukaryotic cells. Correct balance between AS isoforms takes part in molecular mechanisms that properly define spatiotemporal and tissue specific transcriptional programs in physiological conditions. However, several diseases are associated to or even caused by AS alterations. In particular, multiple AS changes occur in cancer cells and sustain the oncogenic transcriptional program. Transcription factors (TFs) represent a key class of proteins that control gene expression by direct binding to DNA regulatory elements. AS events can generate cancer-associated TF isoforms with altered activity, leading to sustained proliferative signaling, differentiation block and apoptosis resistance, all well-known hallmarks of cancer. In this review, we focus on how AS can produce TFs isoforms with opposite transcriptional activities or antagonistic functions that severely impact on cancer biology. This summary points the attention to the relevance of the analysis of TFs splice variants in cancer, which can allow patients stratification despite the presence of interindividual genetic heterogeneity. Recurrent TFs variants that give advantage to specific cancer types not only open the opportunity to use AS transcripts as clinical biomarkers but also guide the development of new anti-cancer strategies in personalized medicine.
Keywords: alternative splicing; cancer; cell death; cell differentiation; cell proliferation; transcription factors.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

