Connexin/Innexin Channels in Cytoplasmic Organelles. Are There Intracellular Gap Junctions? A Hypothesis!
- PMID: 32245189
- PMCID: PMC7139775
- DOI: 10.3390/ijms21062163
Connexin/Innexin Channels in Cytoplasmic Organelles. Are There Intracellular Gap Junctions? A Hypothesis!
Abstract
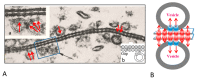
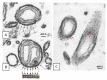
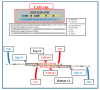
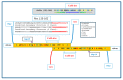
This paper proposes the hypothesis that cytoplasmic organelles directly interact with each other and with gap junctions forming intracellular junctions. This hypothesis originated over four decades ago based on the observation that vesicles lining gap junctions of crayfish giant axons contain electron-opaque particles, similar in size to junctional innexons that often appear to directly interact with junctional innexons; similar particles were seen also in the outer membrane of crayfish mitochondria. Indeed, vertebrate connexins assembled into hexameric connexons are present not only in the membranes of the Golgi apparatus but also in those of the mitochondria and endoplasmic reticulum. It seems possible, therefore, that cytoplasmic organelles may be able to exchange small molecules with each other as well as with organelles of coupled cells via gap junctions.
Keywords: calmodulin; cell-to-cell channels; channel gating; connexin; crayfish giant axons; gap junctions; innexin; liver; mitochondria; stomach.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Assembly of heteromeric connexons in guinea-pig liver en route to the Golgi apparatus, plasma membrane and gap junctions.Eur J Biochem. 1999 May;262(1):142-8. doi: 10.1046/j.1432-1327.1999.00343.x. Eur J Biochem. 1999. PMID: 10231375
-
Calmodulin-Connexin Partnership in Gap Junction Channel Regulation-Calmodulin-Cork Gating Model.Int J Mol Sci. 2021 Dec 2;22(23):13055. doi: 10.3390/ijms222313055. Int J Mol Sci. 2021. PMID: 34884859 Free PMC article. Review.
-
Analysis of connexin intracellular transport and assembly.Methods. 2000 Feb;20(2):156-64. doi: 10.1006/meth.1999.0933. Methods. 2000. PMID: 10671309
-
Incorporation of connexins into plasma membranes and gap junctions.Cardiovasc Res. 2004 May 1;62(2):378-87. doi: 10.1016/j.cardiores.2004.01.016. Cardiovasc Res. 2004. PMID: 15094357 Review.
-
Emerging issues of connexin channels: biophysics fills the gap.Q Rev Biophys. 2001 Aug;34(3):325-472. doi: 10.1017/s0033583501003705. Q Rev Biophys. 2001. PMID: 11838236 Review.
Cited by
-
Calcium Role in Gap Junction Channel Gating: Direct Electrostatic or Calmodulin-Mediated?Int J Mol Sci. 2024 Sep 10;25(18):9789. doi: 10.3390/ijms25189789. Int J Mol Sci. 2024. PMID: 39337278 Free PMC article. Review.
-
The Multifaceted Role of Connexins in Tumor Microenvironment Initiation and Maintenance.Biology (Basel). 2023 Jan 28;12(2):204. doi: 10.3390/biology12020204. Biology (Basel). 2023. PMID: 36829482 Free PMC article. Review.
-
Connexin 43 Role in Mitochondrial Transfer and Homeostasis in the Central Nervous System.J Cell Physiol. 2025 Aug;240(8):e70086. doi: 10.1002/jcp.70086. J Cell Physiol. 2025. PMID: 40838510 Free PMC article. Review.
-
Potential Role of Fenestrated Septa in Axonal Transport of Golgi Cisternae and Gap Junction Formation/Function.Int J Mol Sci. 2023 Mar 11;24(6):5385. doi: 10.3390/ijms24065385. Int J Mol Sci. 2023. PMID: 36982457 Free PMC article. Review.
-
Dynamics of Endothelial Cell Diversity and Plasticity in Health and Disease.Cells. 2024 Jul 29;13(15):1276. doi: 10.3390/cells13151276. Cells. 2024. PMID: 39120307 Free PMC article. Review.
References
-
- Peracchia C. Gap Junctions—Molecular Basis of Cell Communication in Health and Disease. Volume 49 Academic Press; San Diego, CA, USA: 2000. Current Topics in Membranes.
-
- Harris A.L., Locke D. Connexins—A Guide. Humana Press/Springer; New York, NY, USA: 2009.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

