Systems Analysis of NADH Dehydrogenase Mutants Reveals Flexibility and Limits of Pseudomonas taiwanensis VLB120's Metabolism
- PMID: 32245760
- PMCID: PMC7237778
- DOI: 10.1128/AEM.03038-19
Systems Analysis of NADH Dehydrogenase Mutants Reveals Flexibility and Limits of Pseudomonas taiwanensis VLB120's Metabolism
Abstract
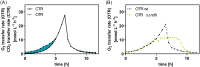
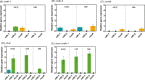
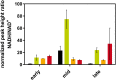
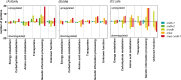
Obligate aerobic organisms rely on a functional electron transport chain for energy conservation and NADH oxidation. Because of this essential requirement, the genes of this pathway are likely constitutively and highly expressed to avoid a cofactor imbalance and energy shortage under fluctuating environmental conditions. We here investigated the essentiality of the three NADH dehydrogenases of the respiratory chain of the obligate aerobe Pseudomonas taiwanensis VLB120 and the impact of the knockouts of corresponding genes on its physiology and metabolism. While a mutant lacking all three NADH dehydrogenases seemed to be nonviable, the single or double knockout mutant strains displayed no, or only a weak, phenotype. Only the mutant deficient in both type 2 dehydrogenases showed a clear phenotype with biphasic growth behavior and a strongly reduced growth rate in the second phase. In-depth analyses of the metabolism of the generated mutants, including quantitative physiological experiments, transcript analysis, proteomics, and enzyme activity assays revealed distinct responses to type 2 and type 1 dehydrogenase deletions. An overall high metabolic flexibility enables P. taiwanensis to cope with the introduced genetic perturbations and maintain stable phenotypes, likely by rerouting of metabolic fluxes. This metabolic adaptability has implications for biotechnological applications. While the phenotypic robustness is favorable in large-scale applications with inhomogeneous conditions, the possible versatile redirecting of carbon fluxes upon genetic interventions can thwart metabolic engineering efforts.IMPORTANCE While Pseudomonas has the capability for high metabolic activity and the provision of reduced redox cofactors important for biocatalytic applications, exploitation of this characteristic might be hindered by high, constitutive activity of and, consequently, competition with the NADH dehydrogenases of the respiratory chain. The in-depth analysis of NADH dehydrogenase mutants of Pseudomonas taiwanensis VLB120 presented here provides insight into the phenotypic and metabolic response of this strain to these redox metabolism perturbations. This high degree of metabolic flexibility needs to be taken into account for rational engineering of this promising biotechnological workhorse toward a host with a controlled and efficient supply of redox cofactors for product synthesis.
Keywords: NADH dehydrogenase; Pseudomonas; electron transport chain; oxidative stress; pseudomonads; redox metabolism; respiratory activity.
Copyright © 2020 American Society for Microbiology.
Figures

Similar articles
-
Shewanella oneidensis MR-1 Utilizes both Sodium- and Proton-Pumping NADH Dehydrogenases during Aerobic Growth.Appl Environ Microbiol. 2018 May 31;84(12):e00415-18. doi: 10.1128/AEM.00415-18. Print 2018 Jun 15. Appl Environ Microbiol. 2018. PMID: 29654176 Free PMC article.
-
The three NADH dehydrogenases of Pseudomonas aeruginosa: Their roles in energy metabolism and links to virulence.PLoS One. 2021 Feb 3;16(2):e0244142. doi: 10.1371/journal.pone.0244142. eCollection 2021. PLoS One. 2021. PMID: 33534802 Free PMC article.
-
Constitutively solvent-tolerant Pseudomonas taiwanensis VLB120∆ C∆ ttgV supports particularly high-styrene epoxidation activities when grown under glucose excess conditions.Biotechnol Bioeng. 2019 May;116(5):1089-1101. doi: 10.1002/bit.26924. Epub 2019 Feb 6. Biotechnol Bioeng. 2019. PMID: 30636283
-
Redox rebalance against genetic perturbations and modulation of central carbon metabolism by the oxidative stress regulation.Biotechnol Adv. 2019 Dec;37(8):107441. doi: 10.1016/j.biotechadv.2019.107441. Epub 2019 Aug 28. Biotechnol Adv. 2019. PMID: 31472206 Review.
-
Organization and regulation of the cytosolic NADH metabolism in the yeast Saccharomyces cerevisiae.Mol Cell Biochem. 2004 Jan-Feb;256-257(1-2):73-81. doi: 10.1023/b:mcbi.0000009888.79484.fd. Mol Cell Biochem. 2004. PMID: 14977171 Review.
Cited by
-
Cytochrome c Reductase is a Key Enzyme Involved in the Extracellular Electron Transfer Pathway towards Transition Metal Complexes in Pseudomonas Putida.ChemSusChem. 2020 Oct 7;13(19):5308-5317. doi: 10.1002/cssc.202001645. Epub 2020 Aug 17. ChemSusChem. 2020. PMID: 32678505 Free PMC article.
-
Crp and Arc system directly regulate the transcription of NADH dehydrogenase genes in Shewanella oneidensis nitrate and nitrite respiration.Microbiol Spectr. 2025 Jul;13(7):e0332424. doi: 10.1128/spectrum.03324-24. Epub 2025 May 16. Microbiol Spectr. 2025. PMID: 40377311 Free PMC article.
-
The energy-converting hydrogenase Ech2 is important for the growth of the thermophilic acetogen Thermoanaerobacter kivui on ferredoxin-dependent substrates.Microbiol Spectr. 2024 Apr 2;12(4):e0338023. doi: 10.1128/spectrum.03380-23. Epub 2024 Feb 22. Microbiol Spectr. 2024. PMID: 38385688 Free PMC article.
References
-
- Yim H, Haselbeck R, Niu W, Pujol-Baxley C, Burgard A, Boldt J, Khandurina J, Trawick JD, Osterhout RE, Stephen R, Estadilla J, Teisan S, Schreyer HB, Andrae S, Yang TH, Lee SY, Burk MJ, Van Dien S. 2011. Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol. Nat Chem Biol 7:445–452. doi:10.1038/nchembio.580. - DOI - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

