Binding of Phage-Encoded FlaGrab to Motile Campylobacter jejuni Flagella Inhibits Growth, Downregulates Energy Metabolism, and Requires Specific Flagellar Glycans
- PMID: 32265863
- PMCID: PMC7099621
- DOI: 10.3389/fmicb.2020.00397
Binding of Phage-Encoded FlaGrab to Motile Campylobacter jejuni Flagella Inhibits Growth, Downregulates Energy Metabolism, and Requires Specific Flagellar Glycans
Abstract
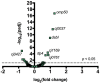
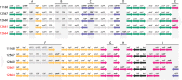
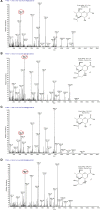
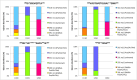
Many bacterial pathogens display glycosylated surface structures that contribute to virulence, and targeting these structures is a viable strategy for pathogen control. The foodborne pathogen Campylobacter jejuni expresses a vast diversity of flagellar glycans, and flagellar glycosylation is essential for its virulence. Little is known about why C. jejuni encodes such a diverse set of flagellar glycans, but it has been hypothesized that evolutionary pressure from bacteriophages (phages) may have contributed to this diversity. However, interactions between Campylobacter phages and host flagellar glycans have not been characterized in detail. Previously, we observed that Gp047 (now renamed FlaGrab), a conserved Campylobacter phage protein, binds to C. jejuni flagella displaying the nine-carbon monosaccharide 7-acetamidino-pseudaminic acid, and that this binding partially inhibits cell growth. However, the mechanism of this growth inhibition, as well as how C. jejuni might resist this activity, are not well-understood. Here we use RNA-Seq to show that FlaGrab exposure leads C. jejuni 11168 cells to downregulate expression of energy metabolism genes, and that FlaGrab-induced growth inhibition is dependent on motile flagella. Our results are consistent with a model whereby FlaGrab binding transmits a signal through flagella that leads to retarded cell growth. To evaluate mechanisms of FlaGrab resistance in C. jejuni, we characterized the flagellar glycans and flagellar glycosylation loci of two C. jejuni strains naturally resistant to FlaGrab binding. Our results point toward flagellar glycan diversity as the mechanism of resistance to FlaGrab. Overall, we have further characterized the interaction between this phage-encoded flagellar glycan-binding protein and C. jejuni, both in terms of mechanism of action and mechanism of resistance. Our results suggest that C. jejuni encodes as-yet unidentified mechanisms for generating flagellar glycan diversity, and point to phage proteins as exciting lenses through which to study bacterial surface glycans.
Keywords: Campylobacter jejuni; bacterial surface structures; bacteriophages; flagella; mass spectrometry; motility; protein glycosylation; pseudaminic acid.
Copyright © 2020 Sacher, Shajahan, Butcher, Patry, Flint, Hendrixson, Stintzi, Azadi and Szymanski.
Figures



Similar articles
-
A Flagellar Glycan-Specific Protein Encoded by Campylobacter Phages Inhibits Host Cell Growth.Viruses. 2015 Dec 16;7(12):6661-74. doi: 10.3390/v7122964. Viruses. 2015. PMID: 26694450 Free PMC article.
-
A receptor-binding protein of Campylobacter jejuni bacteriophage NCTC 12673 recognizes flagellin glycosylated with acetamidino-modified pseudaminic acid.Mol Microbiol. 2015 Jan;95(1):101-15. doi: 10.1111/mmi.12849. Epub 2014 Nov 21. Mol Microbiol. 2015. PMID: 25354466
-
Reduced Infection Efficiency of Phage NCTC 12673 on Non-Motile Campylobacter jejuni Strains Is Related to Oxidative Stress.Viruses. 2021 Sep 29;13(10):1955. doi: 10.3390/v13101955. Viruses. 2021. PMID: 34696385 Free PMC article.
-
Campylobacter flagella: not just for motility.Trends Microbiol. 2007 Oct;15(10):456-61. doi: 10.1016/j.tim.2007.09.006. Epub 2007 Oct 24. Trends Microbiol. 2007. PMID: 17920274 Review.
-
Identifying the targets and functions of N-linked protein glycosylation in Campylobacter jejuni.Mol Omics. 2020 Aug 10;16(4):287-304. doi: 10.1039/d0mo00032a. Mol Omics. 2020. PMID: 32347268 Review.
Cited by
-
Identification of Receptor Binding Proteins in Flagellotropic Agrobacterium Phage 7-7-1.Viruses. 2021 Jun 29;13(7):1267. doi: 10.3390/v13071267. Viruses. 2021. PMID: 34209785 Free PMC article.
-
OmpC and OmpF Outer Membrane Proteins of Escherichia coli and Salmonella enterica Form Bona Fide Amyloids.Int J Mol Sci. 2023 Oct 24;24(21):15522. doi: 10.3390/ijms242115522. Int J Mol Sci. 2023. PMID: 37958507 Free PMC article.
-
RopB protein of Rhizobium leguminosarum bv. viciae adopts amyloid state during symbiotic interactions with pea (Pisum sativum L.).Front Plant Sci. 2022 Oct 24;13:1014699. doi: 10.3389/fpls.2022.1014699. eCollection 2022. Front Plant Sci. 2022. PMID: 36388578 Free PMC article.
-
Bacterial glycosylation, it's complicated.Front Mol Biosci. 2022 Sep 30;9:1015771. doi: 10.3389/fmolb.2022.1015771. eCollection 2022. Front Mol Biosci. 2022. PMID: 36250013 Free PMC article. Review.
-
Campylobacter jejuni Developed the Resistance to Bacteriophage CP39 by Phase Variable Expression of 06875 Encoding the CGPTase.Viruses. 2022 Feb 26;14(3):485. doi: 10.3390/v14030485. Viruses. 2022. PMID: 35336892 Free PMC article.
References
-
- Amour C., Gratz J., Mduma E., Svensen E., Rogawski E. T., McGrath M., et al. . (2016). Epidemiology and impact of campylobacter infection in children in 8 low-resource settings: results from the MAL-ED study. Clin. Infect. Diseases: An Off. Publ. Infect. Diseases Soc. Am. 63, 1171–1179. 10.1093/cid/ciw542 - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

