Heterologous Expression of the AtNPR1 Gene in Olive and Its Effects on Fungal Tolerance
- PMID: 32265961
- PMCID: PMC7100536
- DOI: 10.3389/fpls.2020.00308
Heterologous Expression of the AtNPR1 Gene in Olive and Its Effects on Fungal Tolerance
Abstract
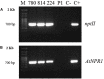
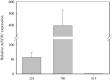
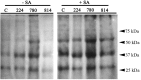
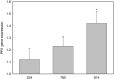
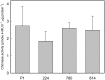
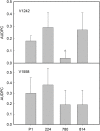
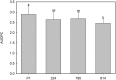
The NPR1 gene encodes a key component of systemic acquired resistance (SAR) signaling mediated by salicylic acid (SA). Overexpression of NPR1 confers resistance to biotrophic and hemibiotrophic fungi in several plant species. The NPR1 gene has also been shown to be involved in the crosstalk between SAR signaling and the jasmonic acid-ethylene (JA/Et) pathway, which is involved in the response to necrotrophic fungi. The aim of this research was to generate transgenic olive plants expressing the NPR1 gene from Arabidopsis thaliana to evaluate their differential response to the hemibiotrophic fungus Verticillium dahliae and the necrotroph Rosellinia necatrix. Three transgenic lines expressing the AtNPR1 gene under the control of the constitutive promoter CaMV35S were obtained using an embryogenic line derived from a seed of cv. Picual. After maturation and germination of the transgenic somatic embryos, the plants were micropropagated and acclimated to ex vitro conditions. The level of AtNPR1 expression in the transgenic materials varied greatly among the different lines and was higher in the NPR1-780 line. The expression of AtNPR1 did not alter the growth of transgenic plants either in vitro or in the greenhouse. Different levels of transgene expression also did not affect basal endochitinase activity in the leaves, which was similar to that of control plants. Response to the hemibiotrophic pathogen V. dahliae varied with pathotype. All plants died by 50 days after inoculation with defoliating (D) pathotype V-138, but the response to non-defoliating (ND) strains differed by race: following inoculation with the V-1242 strain (ND, race 2), symptoms appeared after 44-55 days, with line NPR1-780 showing the lowest disease severity index. This line also showed good performance when inoculated with the V-1558 strain (ND, race 1), although the differences from the control were not statistically significant. In response to the necrotroph R. necatrix, all the transgenic lines showed a slight delay in disease development, with mean area under the disease progress curve (AUDPC) values 7-15% lower than that of the control.
Keywords: Olea europaea; SAR response; Verticillium wilt; genetic transformation; soil-borne pathogens; white root rot.
Copyright © 2020 Narváez, Pliego Prieto, Palomo-Ríos, Fresta, Jiménez-Díaz, Trapero-Casas, Lopez-Herrera, Arjona-Lopez, Mercado and Pliego-Alfaro.
Figures

References
-
- Campbell C. L., Madden L. V. (1990). Introduction to Plant Disease Epidemiology. New York, NY: John Wiley and Sons Inc.
LinkOut - more resources
Full Text Sources
Miscellaneous

