Selective targeting of mutually exclusive DNA G-quadruplexes: HIV-1 LTR as paradigmatic model
- PMID: 32282912
- PMCID: PMC7229848
- DOI: 10.1093/nar/gkaa186
Selective targeting of mutually exclusive DNA G-quadruplexes: HIV-1 LTR as paradigmatic model
Abstract
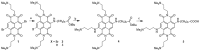
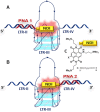
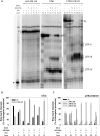
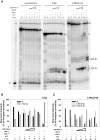
Targeting of G-quadruplexes, non-canonical conformations that form in G-rich regions of nucleic acids, has been proposed as a novel therapeutic strategy toward several diseases, including cancer and infections. The unavailability of highly selective molecules targeting a G-quadruplex of choice has hampered relevant applications. Herein, we describe a novel approach, based on naphthalene diimide (NDI)-peptide nucleic acid (PNA) conjugates, taking advantage of the cooperative interaction of the NDI with the G-quadruplex structure and hybridization of the PNA with the flanking region upstream or downstream the targeted G-quadruplex. By biophysical and biomolecular assays, we show that the NDI-PNA conjugates are able to specifically recognize the G-quadruplex of choice within the HIV-1 LTR region, consisting of overlapping and therefore mutually exclusive G-quadruplexes. Additionally, the conjugates can induce and stabilize the least populated G-quadruplex at the expenses of the more stable ones. The general and straightforward design and synthesis, which readily apply to any G4 target of choice, together with both the red-fluorescent emission and the possibility to introduce cellular localization signals, make the novel conjugates available to selectively control G-quadruplex folding over a wide range of applications.
© The Author(s) 2020. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



Similar articles
-
Synthesis, Binding and Antiviral Properties of Potent Core-Extended Naphthalene Diimides Targeting the HIV-1 Long Terminal Repeat Promoter G-Quadruplexes.J Med Chem. 2015 Dec 24;58(24):9639-52. doi: 10.1021/acs.jmedchem.5b01283. Epub 2015 Dec 8. J Med Chem. 2015. PMID: 26599611 Free PMC article.
-
Naphthalene Diimide-Tetraazacycloalkane Conjugates Are G-Quadruplex-Based HIV-1 Inhibitors with a Dual Mode of Action.ACS Infect Dis. 2024 Feb 9;10(2):489-499. doi: 10.1021/acsinfecdis.3c00453. Epub 2024 Jan 4. ACS Infect Dis. 2024. PMID: 38175706 Free PMC article.
-
DNA Binding Mode Analysis of a Core-Extended Naphthalene Diimide as a Conformation-Sensitive Fluorescent Probe of G-Quadruplex Structures.Int J Mol Sci. 2021 Sep 30;22(19):10624. doi: 10.3390/ijms221910624. Int J Mol Sci. 2021. PMID: 34638964 Free PMC article.
-
Naphthalene Diimides as Multimodal G-Quadruplex-Selective Ligands.Molecules. 2019 Jan 24;24(3):426. doi: 10.3390/molecules24030426. Molecules. 2019. PMID: 30682828 Free PMC article. Review.
-
Targeting DNA G-quadruplex structures with peptide nucleic acids.Curr Pharm Des. 2012;18(14):1984-91. doi: 10.2174/138161212799958440. Curr Pharm Des. 2012. PMID: 22376112 Free PMC article. Review.
Cited by
-
The effect of hairpin loop on the structure and gene expression activity of the long-loop G-quadruplex.Nucleic Acids Res. 2021 Oct 11;49(18):10689-10706. doi: 10.1093/nar/gkab739. Nucleic Acids Res. 2021. PMID: 34450640 Free PMC article.
-
G4-Ligand-Conjugated Oligonucleotides Mediate Selective Binding and Stabilization of Individual G4 DNA Structures.J Am Chem Soc. 2024 Mar 13;146(10):6926-6935. doi: 10.1021/jacs.3c14408. Epub 2024 Mar 2. J Am Chem Soc. 2024. PMID: 38430200 Free PMC article.
-
G-Quadruplex Targeting in the Fight against Viruses: An Update.Int J Mol Sci. 2021 Oct 12;22(20):10984. doi: 10.3390/ijms222010984. Int J Mol Sci. 2021. PMID: 34681641 Free PMC article. Review.
-
Selective Recognition of a Single HIV-1 G-Quadruplex by Ultrafast Small-Molecule Screening.Anal Chem. 2021 Nov 23;93(46):15243-15252. doi: 10.1021/acs.analchem.0c04106. Epub 2021 Nov 11. Anal Chem. 2021. PMID: 34762806 Free PMC article.
-
Light-Activated Molecules Targeting G-Quadruplex Nucleic Acids.Chemistry. 2025 Jul 11;31(39):e202501545. doi: 10.1002/chem.202501545. Epub 2025 Jun 23. Chemistry. 2025. PMID: 40504571 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

