Viral zoonotic risk is homogenous among taxonomic orders of mammalian and avian reservoir hosts
- PMID: 32284401
- PMCID: PMC7196766
- DOI: 10.1073/pnas.1919176117
Viral zoonotic risk is homogenous among taxonomic orders of mammalian and avian reservoir hosts
Abstract
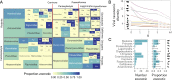
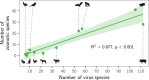
The notion that certain animal groups disproportionately maintain and transmit viruses to humans due to broad-scale differences in ecology, life history, and physiology currently influences global health surveillance and research in disease ecology, virology, and immunology. To directly test whether such "special reservoirs" of zoonoses exist, we used literature searches to construct the largest existing dataset of virus-reservoir relationships, consisting of the avian and mammalian reservoir hosts of 415 RNA and DNA viruses along with their histories of human infection. Reservoir host effects on the propensity of viruses to have been reported as infecting humans were rare and when present were restricted to one or two viral families. The data instead support a largely host-neutral explanation for the distribution of human-infecting viruses across the animal orders studied. After controlling for higher baseline viral richness in mammals versus birds, the observed number of zoonoses per animal order increased as a function of their species richness. Animal orders of established importance as zoonotic reservoirs including bats and rodents were unexceptional, maintaining numbers of zoonoses that closely matched expectations for mammalian groups of their size. Our findings show that variation in the frequency of zoonoses among animal orders can be explained without invoking special ecological or immunological relationships between hosts and viruses, pointing to a need to reconsider current approaches aimed at finding and predicting novel zoonoses.
Keywords: generalized additive model; infectious disease; reservoir; surveillance.
Copyright © 2020 the Author(s). Published by PNAS.
Conflict of interest statement
The authors declare no competing interest.
Figures


References
-
- King D. A., Peckham C., Waage J. K., Brownlie J., Woolhouse M. E. J., Infectious diseases: Preparing for the future. Science 313, 1392–1393 (2006). - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

