Dimerization of mitophagy receptor BNIP3L/NIX is essential for recruitment of autophagic machinery
- PMID: 32286918
- PMCID: PMC8143235
- DOI: 10.1080/15548627.2020.1755120
Dimerization of mitophagy receptor BNIP3L/NIX is essential for recruitment of autophagic machinery
Abstract
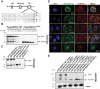
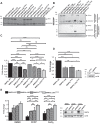
Mitophagy is a conserved intracellular catabolic process responsible for the selective removal of dysfunctional or superfluous mitochondria to maintain mitochondrial quality and need in cells. Here, we examine the mechanisms of receptor-mediated mitophagy activation, with the focus on BNIP3L/NIX mitophagy receptor, proven to be indispensable for selective removal of mitochondria during the terminal differentiation of reticulocytes. The molecular mechanisms of selecting damaged mitochondria from healthy ones are still very obscure. We investigated BNIP3L dimerization as a potentially novel molecular mechanism underlying BNIP3L-dependent mitophagy. Forming stable homodimers, BNIP3L recruits autophagosomes more robustly than its monomeric form. Amino acid substitutions of key transmembrane residues of BNIP3L, BNIP3LG204A or BNIP3LG208V, led to the abolishment of dimer formation, resulting in the lower LC3A-BNIP3L recognition and subsequently lower mitophagy induction. Moreover, we identified the serine 212 as the main amino acid residue at the C-terminal of BNIP3L, which extends to the intermembrane space, responsible for dimerization. In accordance, the phosphomimetic mutation BNIP3LS212E leads to a complete loss of BNIP3L dimerization. Thus, the interplay between BNIP3L phosphorylation and dimerization indicates that the combined mechanism of LIR phosphorylation and receptor dimerization is needed for proper BNIP3L-dependent mitophagy initiation and progression.Abbreviations: AMBRA1: autophagy and beclin 1 regulator 1; Baf A1: bafilomycin A1; BH3: BCL2 homology 3; BNIP3: BCL2 interacting protein 3; BNIP3L/NIX: BCL2 interacting protein 3 like; CCCP: carbonyl cyanide 3-chlorophenylhydrazone; CoCl2: cobalt (II) chloride; FKBP8: FKBP prolyl isomerase 8; FUNDC1: FUN14 domain containing 1; GABARAP: GABA type A receptor-associated protein; GST: glutathione S-transferase; IMM: inner mitochondrial membrane; LIR: LC3-interacting region; MAP1LC3/LC3: microtubule associated protein 1 light chain 3; OMM: outer mitochondrial membrane; PHB2: prohibitin 2; PI: propidium iodide; PINK1: PTEN induced kinase 1; TM: transmembrane domain; TOMM20: translocase of outer mitochondrial membrane 20.
Keywords: Autophagy; BNIP3L/NIX; dimerization; mitophagy; selective autophagy.
Conflict of interest statement
The authors declare no potential conflict of interest.
Figures


Similar articles
-
A brief overview of BNIP3L/NIX receptor-mediated mitophagy.FEBS Open Bio. 2021 Dec;11(12):3230-3236. doi: 10.1002/2211-5463.13307. Epub 2021 Oct 11. FEBS Open Bio. 2021. PMID: 34597467 Free PMC article. Review.
-
Mitochondria ROS and mitophagy in acute kidney injury.Autophagy. 2023 Feb;19(2):401-414. doi: 10.1080/15548627.2022.2084862. Epub 2022 Jun 9. Autophagy. 2023. PMID: 35678504 Free PMC article.
-
Clearance of damaged mitochondria via mitophagy is important to the protective effect of ischemic preconditioning in kidneys.Autophagy. 2019 Dec;15(12):2142-2162. doi: 10.1080/15548627.2019.1615822. Epub 2019 May 22. Autophagy. 2019. PMID: 31066324 Free PMC article.
-
Emerging views of mitophagy in immunity and autoimmune diseases.Autophagy. 2020 Jan;16(1):3-17. doi: 10.1080/15548627.2019.1603547. Epub 2019 Apr 21. Autophagy. 2020. PMID: 30951392 Free PMC article.
-
Organelle-specific autophagy in inflammatory diseases: a potential therapeutic target underlying the quality control of multiple organelles.Autophagy. 2021 Feb;17(2):385-401. doi: 10.1080/15548627.2020.1725377. Epub 2020 Feb 12. Autophagy. 2021. PMID: 32048886 Free PMC article. Review.
Cited by
-
The ER membrane protein complex restricts mitophagy by controlling BNIP3 turnover.EMBO J. 2024 Jan;43(1):32-60. doi: 10.1038/s44318-023-00006-z. Epub 2023 Dec 15. EMBO J. 2024. PMID: 38177312 Free PMC article.
-
GAK and PRKCD are positive regulators of PRKN-independent mitophagy.Nat Commun. 2021 Oct 20;12(1):6101. doi: 10.1038/s41467-021-26331-7. Nat Commun. 2021. PMID: 34671015 Free PMC article.
-
Nix interacts with WIPI2 to induce mitophagy.EMBO J. 2023 Nov 15;42(22):e113491. doi: 10.15252/embj.2023113491. Epub 2023 Aug 25. EMBO J. 2023. PMID: 37621214 Free PMC article.
-
The multifaceted regulation of mitophagy by endogenous metabolites.Autophagy. 2022 Jun;18(6):1216-1239. doi: 10.1080/15548627.2021.1975914. Epub 2021 Sep 29. Autophagy. 2022. PMID: 34583624 Free PMC article.
-
A brief overview of BNIP3L/NIX receptor-mediated mitophagy.FEBS Open Bio. 2021 Dec;11(12):3230-3236. doi: 10.1002/2211-5463.13307. Epub 2021 Oct 11. FEBS Open Bio. 2021. PMID: 34597467 Free PMC article. Review.
References
-
- Kirkin V, Lamark T, Sou Y-S, et al. A role for NBR1 in autophagosomal degradation of ubiquitinated substrates. Mol Cell. 2009;33(4):505–516. - PubMed
-
- Pankiv S, Clausen TH, Lamark T, et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J. Biol. Chem. 2007. August;282(33):24131–24145. - PubMed
-
- N. von M. & F. R, Thurston TLM, Ryzhakov G, Bloor S.. The TBK1 adaptor and autophagy receptor NDP52 restricts the proliferation of ubiquitin-coated bacteria. Nat Immunol. 2009;10(11):1215–1222. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
