Loss of the neural-specific BAF subunit ACTL6B relieves repression of early response genes and causes recessive autism
- PMID: 32312822
- PMCID: PMC7211998
- DOI: 10.1073/pnas.1908238117
Loss of the neural-specific BAF subunit ACTL6B relieves repression of early response genes and causes recessive autism
Abstract
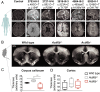
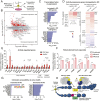
Synaptic activity in neurons leads to the rapid activation of genes involved in mammalian behavior. ATP-dependent chromatin remodelers such as the BAF complex contribute to these responses and are generally thought to activate transcription. However, the mechanisms keeping such "early activation" genes silent have been a mystery. In the course of investigating Mendelian recessive autism, we identified six families with segregating loss-of-function mutations in the neuronal BAF (nBAF) subunit ACTL6B (originally named BAF53b). Accordingly, ACTL6B was the most significantly mutated gene in the Simons Recessive Autism Cohort. At least 14 subunits of the nBAF complex are mutated in autism, collectively making it a major contributor to autism spectrum disorder (ASD). Patient mutations destabilized ACTL6B protein in neurons and rerouted dendrites to the wrong glomerulus in the fly olfactory system. Humans and mice lacking ACTL6B showed corpus callosum hypoplasia, indicating a conserved role for ACTL6B in facilitating neural connectivity. Actl6b knockout mice on two genetic backgrounds exhibited ASD-related behaviors, including social and memory impairments, repetitive behaviors, and hyperactivity. Surprisingly, mutation of Actl6b relieved repression of early response genes including AP1 transcription factors (Fos, Fosl2, Fosb, and Junb), increased chromatin accessibility at AP1 binding sites, and transcriptional changes in late response genes associated with early response transcription factor activity. ACTL6B loss is thus an important cause of recessive ASD, with impaired neuron-specific chromatin repression indicated as a potential mechanism.
Keywords: BAF; activity dependent; autism; mouse model; recessive.
Copyright © 2020 the Author(s). Published by PNAS.
Conflict of interest statement
Competing interest statement: G.R.C. is a founder of Foghorn Therapeutics.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
- F32 MH103949/MH/NIMH NIH HHS/United States
- F99 NS118735/NS/NINDS NIH HHS/United States
- U54 HG003067/HG/NHGRI NIH HHS/United States
- R01 NS046789/NS/NINDS NIH HHS/United States
- UL1 TR001863/TR/NCATS NIH HHS/United States
- T32 GM145427/GM/NIGMS NIH HHS/United States
- R37 NS046789/NS/NINDS NIH HHS/United States
- HHMI/Howard Hughes Medical Institute/United States
- F31 MH116588/MH/NIMH NIH HHS/United States
- R01 NS048453/NS/NINDS NIH HHS/United States
- P50 DA042012/DA/NIDA NIH HHS/United States
- U54 HG006504/HG/NHGRI NIH HHS/United States
- R01 CA163915/CA/NCI NIH HHS/United States
- T32 GM008666/GM/NIGMS NIH HHS/United States
- T32 GM007790/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

