ShallowHRD: detection of homologous recombination deficiency from shallow whole genome sequencing
- PMID: 32315385
- PMCID: PMC7320600
- DOI: 10.1093/bioinformatics/btaa261
ShallowHRD: detection of homologous recombination deficiency from shallow whole genome sequencing
Abstract
Summary: We introduce shallowHRD, a software tool to evaluate tumor homologous recombination deficiency (HRD) based on whole genome sequencing (WGS) at low coverage (shallow WGS or sWGS; ∼1X coverage). The tool, based on mining copy number alterations profile, implements a fast and straightforward procedure that shows 87.5% sensitivity and 90.5% specificity for HRD detection. shallowHRD could be instrumental in predicting response to poly(ADP-ribose) polymerase inhibitors, to which HRD tumors are selectively sensitive. shallowHRD displays efficiency comparable to most state-of-art approaches, is cost-effective, generates low-storable outputs and is also suitable for fixed-formalin paraffin embedded tissues.
Availability and implementation: shallowHRD R script and documentation are available at https://github.com/aeeckhou/shallowHRD.
Supplementary information: Supplementary data are available at Bioinformatics online.
© The Author(s) 2020. Published by Oxford University Press.
Figures
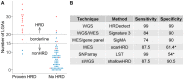
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

