Rice GERMIN-LIKE PROTEIN 2-1 Functions in Seed Dormancy under the Control of Abscisic Acid and Gibberellic Acid Signaling Pathways
- PMID: 32321839
- PMCID: PMC7333727
- DOI: 10.1104/pp.20.00253
Rice GERMIN-LIKE PROTEIN 2-1 Functions in Seed Dormancy under the Control of Abscisic Acid and Gibberellic Acid Signaling Pathways
Abstract
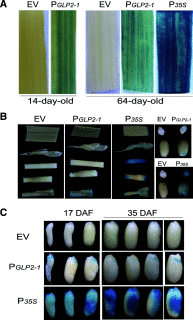
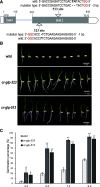
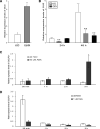
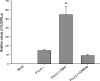
Seed dormancy is a natural phenomenon in plants. It ensures that seeds complete the grain-filling stage before germination and prevents germination in unsuitable ecological conditions. In this study, we determined the previously unknown function of the rice (Oryza sativa) gene GERMIN-LIKE PROTEIN 2-1 (OsGLP2-1) in seed dormancy. Using artificial microRNA and CRISPR/CAS9 approaches, suppression of OsGLP2-1 expression in rice resulted in the release of dormancy in immature seeds. Conversely, overexpression of OsGLP2-1 driven by the OsGLP2-1 native promoter led to greater seed dormancy. Seed scutellum-specific expression of OsGLP2-1 was increased by exogenous abscisic acid, but decreased with gibberellic acid treatment. We provide evidence that OsGLP2-1 is antagonistically controlled at the transcriptional level by ABA INSENSITIVE5 and GAMYB transcription factors. We conclude that OsGLP2-1 acts as a buffer, maintaining appropriate equilibrium for the regulation of primary dormancy during seed development in rice.
© 2020 American Society of Plant Biologists. All Rights Reserved.
Figures

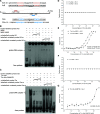

References
-
- Alboresi A, Gestin C, Leydecker MT, Bedu M, Meyer C, Truong HN(2005) Nitrate, a signal relieving seed dormancy in Arabidopsis. Plant Cell Environ 28: 500–512 - PubMed
-
- Asakura T, Hirose S, Asatsuma S, Nanjo Y, Nakaizumi T, Itoh K, Hori H, Komatsu S, Mitsui T(2007) Proteomic characterization of tissue expansion of rice scutellum stimulated by abscisic acid. Biosci Biotechnol Biochem 71: 1260–1268 - PubMed
-
- Berna A, Bernier F(1997) Regulated expression of a wheat germin gene in tobacco: Oxalate oxidase activity and apoplastic localization of the heterologous protein. Plant Mol Biol 33: 417–429 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

