The Flemmingsome reveals an ESCRT-to-membrane coupling via ALIX/syntenin/syndecan-4 required for completion of cytokinesis
- PMID: 32321914
- PMCID: PMC7176721
- DOI: 10.1038/s41467-020-15205-z
The Flemmingsome reveals an ESCRT-to-membrane coupling via ALIX/syntenin/syndecan-4 required for completion of cytokinesis
Abstract
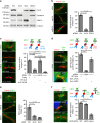
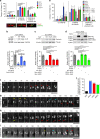
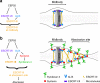
Cytokinesis requires the constriction of ESCRT-III filaments on the side of the midbody, where abscission occurs. After ESCRT recruitment at the midbody, it is not known how the ESCRT-III machinery localizes to the abscission site. To reveal actors involved in abscission, we obtained the proteome of intact, post-abscission midbodies (Flemmingsome) and identified 489 proteins enriched in this organelle. Among these proteins, we further characterized a plasma membrane-to-ESCRT module composed of the transmembrane proteoglycan syndecan-4, ALIX and syntenin, a protein that bridges ESCRT-III/ALIX to syndecans. The three proteins are highly recruited first at the midbody then at the abscission site, and their depletion delays abscission. Mechanistically, direct interactions between ALIX, syntenin and syndecan-4 are essential for proper enrichment of the ESCRT-III machinery at the abscission site, but not at the midbody. We propose that the ESCRT-III machinery must be physically coupled to a membrane protein at the cytokinetic abscission site for efficient scission, uncovering common requirements in cytokinesis, exosome formation and HIV budding.
Conflict of interest statement
The authors declare no competing interests.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

