Multiplex T Cell Stimulation Assay Utilizing a T Cell Activation Reporter-Based Detection System
- PMID: 32328071
- PMCID: PMC7160884
- DOI: 10.3389/fimmu.2020.00633
Multiplex T Cell Stimulation Assay Utilizing a T Cell Activation Reporter-Based Detection System
Abstract
Recent advancements in single cell sequencing technologies allow for identification of numerous immune-receptors expressed by T cells such as tumor-specific and autoimmune T cells. Determining antigen specificity of those cells holds immense therapeutic promise. Therefore, the purpose of this study was to develop a method that can efficiently test antigen reactivity of multiple T cell receptors (TCRs) with limited cost, time, and labor. Nuclear factor of activated T cells (NFAT) is a transcription factor involved in producing cytokines and is often utilized as a reporter system for T cell activation. Using a NFAT-based fluorescent reporter system, we generated T-hybridoma cell lines that express intensely fluorescent proteins in response to antigen stimulation and constitutively express additional fluorescent proteins, which serve as identifiers of each T-hybridoma expressing a unique TCR. This allows for the combination of multiple T-hybridoma lines within a single reaction. Sensitivity to stimulation is not decreased by adding fluorescent proteins or multiplexing T cells. In multiplexed reactions, response by one cell line does not induce response in others, thus preserving specificity. This multiplex assay system will be a useful tool for antigen discovery research in a variety of contexts, including using combinatorial peptide libraries to determine T cell epitopes.
Keywords: T cell receptors; antigens; epitopes; multiplex assay; reporter.
Copyright © 2020 Mann, Zhou, Landry, Anderson, Alkanani, Fischer, Peakman, Mallone, Campbell, Michels and Nakayama.
Figures





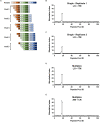

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

