An Improved Phenotype-Driven Tool for Rare Mendelian Variant Prioritization: Benchmarking Exomiser on Real Patient Whole-Exome Data
- PMID: 32340307
- PMCID: PMC7230372
- DOI: 10.3390/genes11040460
An Improved Phenotype-Driven Tool for Rare Mendelian Variant Prioritization: Benchmarking Exomiser on Real Patient Whole-Exome Data
Abstract
Next-generation sequencing has revolutionized rare disease diagnostics, but many patients remain without a molecular diagnosis, particularly because many candidate variants usually survive despite strict filtering. Exomiser was launched in 2014 as a Java tool that performs an integrative analysis of patients' sequencing data and their phenotypes encoded with Human Phenotype Ontology (HPO) terms. It prioritizes variants by leveraging information on variant frequency, predicted pathogenicity, and gene-phenotype associations derived from human diseases, model organisms, and protein-protein interactions. Early published releases of Exomiser were able to prioritize disease-causative variants as top candidates in up to 97% of simulated whole-exomes. The size of the tested real patient datasets published so far are very limited. Here, we present the latest Exomiser version 12.0.1 with many new features. We assessed the performance using a set of 134 whole-exomes from patients with a range of rare retinal diseases and known molecular diagnosis. Using default settings, Exomiser ranked the correct diagnosed variants as the top candidate in 74% of the dataset and top 5 in 94%; not using the patients' HPO profiles (i.e., variant-only analysis) decreased the performance to 3% and 27%, respectively. In conclusion, Exomiser is an effective support tool for rare Mendelian phenotype-driven variant prioritization.
Keywords: bioinformatics; human phenotype ontology; inherited retinal disease; phenotypic similarity; rare disease; variant prioritization; whole-exome sequencing; whole-genome sequencing.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures

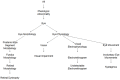
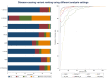

References
-
- Wright C.F., FitzPatrick D.R., Firth H.V. Paediatric genomics: Diagnosing rare disease in children. Nat. Rev. Genet. 2018;19:327. - PubMed

