A Multibasic Cleavage Site in the Spike Protein of SARS-CoV-2 Is Essential for Infection of Human Lung Cells
- PMID: 32362314
- PMCID: PMC7194065
- DOI: 10.1016/j.molcel.2020.04.022
A Multibasic Cleavage Site in the Spike Protein of SARS-CoV-2 Is Essential for Infection of Human Lung Cells
Abstract
The pandemic coronavirus SARS-CoV-2 threatens public health worldwide. The viral spike protein mediates SARS-CoV-2 entry into host cells and harbors a S1/S2 cleavage site containing multiple arginine residues (multibasic) not found in closely related animal coronaviruses. However, the role of this multibasic cleavage site in SARS-CoV-2 infection is unknown. Here, we report that the cellular protease furin cleaves the spike protein at the S1/S2 site and that cleavage is essential for S-protein-mediated cell-cell fusion and entry into human lung cells. Moreover, optimizing the S1/S2 site increased cell-cell, but not virus-cell, fusion, suggesting that the corresponding viral variants might exhibit increased cell-cell spread and potentially altered virulence. Our results suggest that acquisition of a S1/S2 multibasic cleavage site was essential for SARS-CoV-2 infection of humans and identify furin as a potential target for therapeutic intervention.
Keywords: COVID-19; S1/S2; SARS-CoV-2; TMPRSS2; cleavage; entry; furin; membrane fusion; spike.
Copyright © 2020 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of Interest The authors declare not competing interests
Figures

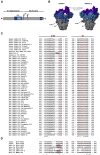

References
-
- Bertram S., Dijkman R., Habjan M., Heurich A., Gierer S., Glowacka I., Welsch K., Winkler M., Schneider H., Hofmann-Winkler H., et al. TMPRSS2 activates the human coronavirus 229E for cathepsin-independent host cell entry and is expressed in viral target cells in the respiratory epithelium. J. Virol. 2013;87:6150–6160. - PMC - PubMed
-
- Gierer S., Bertram S., Kaup F., Wrensch F., Heurich A., Krämer-Kühl A., Welsch K., Winkler M., Meyer B., Drosten C., et al. The spike protein of the emerging betacoronavirus EMC uses a novel coronavirus receptor for entry, can be activated by TMPRSS2, and is targeted by neutralizing antibodies. J. Virol. 2013;87:5502–5511. - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

