Real-time observation of CRISPR spacer acquisition by Cas1-Cas2 integrase
- PMID: 32367067
- PMCID: PMC7241818
- DOI: 10.1038/s41594-020-0415-7
Real-time observation of CRISPR spacer acquisition by Cas1-Cas2 integrase
Abstract
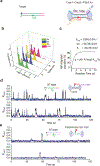
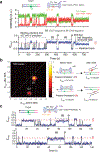
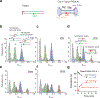
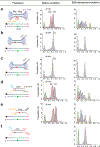
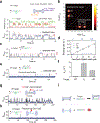
Cas1 integrase associates with Cas2 to insert short DNA fragments into a CRISPR array, establishing nucleic acid memory in prokaryotes. Here we applied single-molecule FRET methods to the Enterococcus faecalis (Efa) Cas1-Cas2 system to establish a kinetic framework describing target-searching, integration, and post-synapsis events. EfaCas1-Cas2 on its own is not able to find the CRISPR repeat in the CRISPR array; it only does so after prespacer loading. The leader sequence adjacent to the repeat further stabilizes EfaCas1-Cas2 contacts, enabling leader-side integration and subsequent spacer-side integration. The resulting post-synaptic complex (PSC) has a surprisingly short mean lifetime. Remarkably, transcription effectively resolves the PSC, and we predict that this is a conserved mechanism that ensures efficient and directional spacer integration in many CRISPR systems. Overall, our study provides a complete model of spacer acquisition, which can be harnessed for DNA-based information storage and cell lineage tracing technologies.
Conflict of interest statement
Competing Interests
The authors declare no competing interests.
Figures












Similar articles
-
How type II CRISPR-Cas establish immunity through Cas1-Cas2-mediated spacer integration.Nature. 2017 Oct 5;550(7674):137-141. doi: 10.1038/nature24020. Epub 2017 Sep 4. Nature. 2017. PMID: 28869593 Free PMC article.
-
Integrase-mediated spacer acquisition during CRISPR-Cas adaptive immunity.Nature. 2015 Mar 12;519(7542):193-8. doi: 10.1038/nature14237. Epub 2015 Feb 18. Nature. 2015. PMID: 25707795 Free PMC article.
-
DNA binding specificities of Escherichia coli Cas1-Cas2 integrase drive its recruitment at the CRISPR locus.Nucleic Acids Res. 2017 Mar 17;45(5):2714-2723. doi: 10.1093/nar/gkw1309. Nucleic Acids Res. 2017. PMID: 28034956 Free PMC article.
-
Harnessing CRISPR-Cas adaptation for RNA recording and beyond.BMB Rep. 2024 Jan;57(1):40-49. doi: 10.5483/BMBRep.2023-0050. BMB Rep. 2024. PMID: 38053290 Free PMC article. Review.
-
Molecular Details of DNA Integration by CRISPR-Associated Proteins During Adaptation in Bacteria and Archaea.Adv Exp Med Biol. 2023;1414:27-43. doi: 10.1007/5584_2022_730. Adv Exp Med Biol. 2023. PMID: 35852729 Review.
Cited by
-
Mechanism for Cas4-assisted directional spacer acquisition in CRISPR-Cas.Nature. 2021 Oct;598(7881):515-520. doi: 10.1038/s41586-021-03951-z. Epub 2021 Sep 29. Nature. 2021. PMID: 34588691 Free PMC article.
-
CRISPR-Associated Primase-Polymerases are implicated in prokaryotic CRISPR-Cas adaptation.Nat Commun. 2021 Jun 17;12(1):3690. doi: 10.1038/s41467-021-23535-9. Nat Commun. 2021. PMID: 34140468 Free PMC article.
-
SspA is a transcriptional regulator of CRISPR adaptation in E. coli.Nucleic Acids Res. 2025 Feb 8;53(4):gkae1244. doi: 10.1093/nar/gkae1244. Nucleic Acids Res. 2025. PMID: 39727179 Free PMC article.
-
Mechanisms used for cDNA synthesis and site-specific integration of RNA into DNA genomes by a reverse transcriptase-Cas1 fusion protein.bioRxiv [Preprint]. 2023 Sep 3:2023.09.01.555893. doi: 10.1101/2023.09.01.555893. bioRxiv. 2023. Update in: Sci Adv. 2024 Apr 12;10(15):eadk8791. doi: 10.1126/sciadv.adk8791. PMID: 37693417 Free PMC article. Updated. Preprint.
-
Disintegration promotes protospacer integration by the Cas1-Cas2 complex.Elife. 2021 Aug 26;10:e65763. doi: 10.7554/eLife.65763. Elife. 2021. PMID: 34435949 Free PMC article.
References
-
- Barrangou R et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007). - PubMed
-
- Makarova KS, Grishin NV, Shabalina SA, Wolf YI & Koonin EV A putative RNA-interference-based immune system in prokaryotes: computational analysis of the predicted enzymatic machinery, functional analogies with eukaryotic RNAi, and hypothetical mechanisms of action. Biol Direct 1, 7, doi:10.1186/1745-6150-1-7 (2006). - DOI - PMC - PubMed
References for the Methods and Supplemental text sections
-
- Joo C & Ha T Labeling Proteins for Single-Molecule FRET. Cold Spring Harbor Protocols 2012, pdb.prot071035 (2012). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

