2.7 Å cryo-EM structure of vitrified M. musculus H-chain apoferritin from a compact 200 keV cryo-microscope
- PMID: 32374767
- PMCID: PMC7202636
- DOI: 10.1371/journal.pone.0232540
2.7 Å cryo-EM structure of vitrified M. musculus H-chain apoferritin from a compact 200 keV cryo-microscope
Abstract
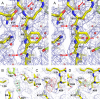
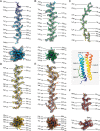
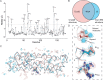
Here we present the structure of mouse H-chain apoferritin at 2.7 Å (FSC = 0.143) solved by single particle cryogenic electron microscopy (cryo-EM) using a 200 kV device, the Thermo Fisher Glacios®. This is a compact, two-lens illumination system with a constant power objective lens, without any energy filters or aberration correctors, often thought of as a "screening cryo-microscope". Coulomb potential maps reveal clear densities for main chain carbonyl oxygens, residue side chains (including alternative conformations) and bound solvent molecules. We used a quasi-crystallographic reciprocal space approach to fit model coordinates to the experimental cryo-EM map. We argue that the advantages offered by (a) the high electronic and mechanical stability of the microscope, (b) the high emission stability and low beam energy spread of the high brightness Field Emission Gun (X-FEG), (c) direct electron detection technology and (d) particle-based Contrast Transfer Function (CTF) refinement have contributed to achieving high resolution. Overall, we show that basic electron optical settings for automated cryo-electron microscopy imaging can be used to determine structures approaching atomic resolution.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

