CRISPR-Cas12a system in fission yeast for multiplex genomic editing and CRISPR interference
- PMID: 32374858
- PMCID: PMC7261154
- DOI: 10.1093/nar/gkaa329
CRISPR-Cas12a system in fission yeast for multiplex genomic editing and CRISPR interference
Abstract
The CRISPR-Cas12a is a class II, type V clustered regularly interspaced short palindromic repeat (CRISPR) system with both RNase and DNase activity. Compared to the CRISPR-Cas9 system, it recognizes T-rich PAM sequences and has the advantage of multiplex genomic editing. Here, in fission yeast Schizosaccharomyces pombe, we successfully implemented the CRISPR-Cas12a system for versatile genomic editing and manipulation. In addition to the rrk1 promoter, we used new pol II promoters from endogenous coding genes to express crRNA for Cas12a and obtained a much higher editing efficiency. This new design expands the promoter choices for potential applications in fission yeast and other organisms. In addition, we expressed a gRNA array using a strong constitutive pol II promoter. The array transcript is processed by Cas12a itself to release multiple mature crRNAs. With this construct, multiplex genomic editing of up to three loci was achieved from a single yeast transformation. We also built a CRISPR interference system using a DNase-dead Cas12a to significantly repress endogenous gene expression. Our study provides the first CRISPR-Cas12a toolkit for efficient and rapid genomic gene editing and regulation in fission yeast.
© The Author(s) 2020. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



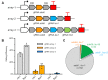

Similar articles
-
Multiplexed conditional genome editing with Cas12a in Drosophila.Proc Natl Acad Sci U S A. 2020 Sep 15;117(37):22890-22899. doi: 10.1073/pnas.2004655117. Epub 2020 Aug 25. Proc Natl Acad Sci U S A. 2020. PMID: 32843348 Free PMC article.
-
Combined genome editing and transcriptional repression for metabolic pathway engineering in Corynebacterium glutamicum using a catalytically active Cas12a.Appl Microbiol Biotechnol. 2019 Nov;103(21-22):8911-8922. doi: 10.1007/s00253-019-10118-4. Epub 2019 Oct 3. Appl Microbiol Biotechnol. 2019. PMID: 31583448
-
Single transcript unit CRISPR 2.0 systems for robust Cas9 and Cas12a mediated plant genome editing.Plant Biotechnol J. 2019 Jul;17(7):1431-1445. doi: 10.1111/pbi.13068. Epub 2019 Jan 17. Plant Biotechnol J. 2019. PMID: 30582653 Free PMC article.
-
CRISPR-Cas12a System for Biosensing and Gene Regulation.Chem Asian J. 2021 Apr 19;16(8):857-867. doi: 10.1002/asia.202100043. Epub 2021 Mar 18. Chem Asian J. 2021. PMID: 33638271 Review.
-
Recent advances of Cas12a applications in bacteria.Appl Microbiol Biotechnol. 2021 Apr;105(8):2981-2990. doi: 10.1007/s00253-021-11243-9. Epub 2021 Mar 23. Appl Microbiol Biotechnol. 2021. PMID: 33754170 Free PMC article. Review.
Cited by
-
CRISPR-Cas Technology for Bioengineering Conventional and Non-Conventional Yeasts: Progress and New Challenges.Int J Mol Sci. 2023 Oct 18;24(20):15310. doi: 10.3390/ijms242015310. Int J Mol Sci. 2023. PMID: 37894990 Free PMC article. Review.
-
Implementation of dCas9-mediated CRISPRi in the fission yeast Schizosaccharomyces pombe.G3 (Bethesda). 2021 Apr 15;11(4):jkab051. doi: 10.1093/g3journal/jkab051. G3 (Bethesda). 2021. PMID: 33617628 Free PMC article.
-
Simplicity within biological complexity.Bioinform Adv. 2025 Feb 6;5(1):vbae164. doi: 10.1093/bioadv/vbae164. eCollection 2025. Bioinform Adv. 2025. PMID: 39927291 Free PMC article.
-
Establishment of the CRISPR-Cpf1 gene editing system in Bacillus licheniformis and multiplexed gene knockout.Synth Syst Biotechnol. 2024 Aug 8;10(1):39-48. doi: 10.1016/j.synbio.2024.08.002. eCollection 2025. Synth Syst Biotechnol. 2024. PMID: 39224148 Free PMC article.
-
The rise and future of CRISPR-based approaches for high-throughput genomics.FEMS Microbiol Rev. 2024 Sep 18;48(5):fuae020. doi: 10.1093/femsre/fuae020. FEMS Microbiol Rev. 2024. PMID: 39085047 Free PMC article. Review.
References
-
- Barrangou R., Fremaux C., Deveau H., Richards M., Boyaval P., Moineau S., Romero D.A., Horvath P.. CRISPR provides acquired resistance against viruses in prokaryotes. Science (New York, N.Y.). 2007; 315:1709–1712. - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

