Ampliconic Genes on the Great Ape Y Chromosomes: Rapid Evolution of Copy Number but Conservation of Expression Levels
- PMID: 32374870
- PMCID: PMC7313670
- DOI: 10.1093/gbe/evaa088
Ampliconic Genes on the Great Ape Y Chromosomes: Rapid Evolution of Copy Number but Conservation of Expression Levels
Abstract
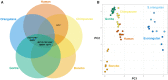
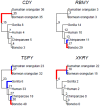
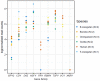
Multicopy ampliconic gene families on the Y chromosome play an important role in spermatogenesis. Thus, studying their genetic variation in endangered great ape species is critical. We estimated the sizes (copy number) of nine Y ampliconic gene families in population samples of chimpanzee, bonobo, and orangutan with droplet digital polymerase chain reaction, combined these estimates with published data for human and gorilla, and produced genome-wide testis gene expression data for great apes. Analyzing this comprehensive data set within an evolutionary framework, we, first, found high inter- and intraspecific variation in gene family size, with larger families exhibiting higher variation as compared with smaller families, a pattern consistent with random genetic drift. Second, for four gene families, we observed significant interspecific size differences, sometimes even between sister species-chimpanzee and bonobo. Third, despite substantial variation in copy number, Y ampliconic gene families' expression levels did not differ significantly among species, suggesting dosage regulation. Fourth, for three gene families, size was positively correlated with gene expression levels across species, suggesting that, given sufficient evolutionary time, copy number influences gene expression. Our results indicate high variability in size but conservation in gene expression levels in Y ampliconic gene families, significantly advancing our understanding of Y-chromosome evolution in great apes.
Keywords: Y chromosome; ampliconic genes; bonobo; gene copy number; gene expression; great apes; orangutan.
© The Author(s) 2020. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures




References
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ.. 1990. Basic local alignment search tool. J Mol Biol. 215(3):403–410. - PubMed
-
- Ancrenaz M, et al. 2016. Pongo pygmaeus. The IUCN Red List of Threatened Species 2016: e.T17975A123809220. Available from: https://dx.doi.org/10.2305/IUCN.UK.2016-1.RLTS.T17975A17966347.en. Accesed May 7, 2020.
-
- Boetzer M, Henkel CV, Jansen HJ, Butler D, Pirovano W.. 2011. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27(4):578–579. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

