[Analysis of variation and evolution of SARS-CoV-2 genome]
- PMID: 32376535
- PMCID: PMC7086142
- DOI: 10.12122/j.issn.1673-4254.2020.02.02
[Analysis of variation and evolution of SARS-CoV-2 genome]
Abstract
Objective: To analyze the evolution and variation of SARS-CoV-2 during the epidemic starting at the end of 2019.
Methods: We downloaded the full-length genome sequence of SARS-CoV-2 from the databases of GISAID and NCBI. Using the software for bioinformatics including MEGA-X, BEAST, and TempEst, we constructed the genomic evolution tree, inferred the time evolution signal of the virus, calculated the tMRCA time of the virus and analyzed the selection pressure of the virus during evolution.
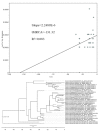
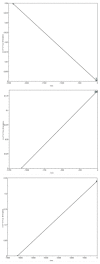
Results: The phylogenetic tree showed that SARS-CoV-2 belonged to the Sarbecovirus subgenus of β Coronavirus genus together with bat coronavirus BetaCoV/bat/Yunnan/RaTG13/2013, bat-SL-CoVZC45, bat-SL-CoVZXC21 and SARS-CoV. The genomic sequences of SARS-CoV-2 isolated from the ongoing epidemic showed a weak time evolution signal with an average tMRCA time of 73 days (95% CI: 38.9-119.3 days). No positive time evolution signal was found between SARS-CoV-2 and BetaCoV/bat/Yunnan/RaTG13/2013, but the former virus had a strong positive temporal evolution relationship with bat-SL-CoVZC45 and SARS-CoV. The major cause for mutations of SARS-CoV-2 was the pressure of purification selection during the epidemic.
Conclusions: SARS-CoV-2 may have emerged as early as November, 2019, originating most likely from bat-associated coronavirus. This finding may provide evidence for tracing the sources and evolution of the virus.
目的: 分析新型冠状病毒SARS-CoV-2的进化、变异情况。
方法: 从GISAID、NCBI中下载相关病毒全基因组序列,运用生物信息学软件MEGA-X、BEAST、TempEst等软件,构建基因组进化树,推测病毒的时间进化信号,计算病毒出现的tMRCA时间,分析病毒进化的选择压力。
结果: 基因组进化树显示SARS-CoV-2与蝙蝠冠状病毒Beta CoV/bat/Yunnan/RaTG13/2013、bat-SL-CoVZC45、bat-SL-CoVZXC21和SARS-CoV等病毒共同构成冠状病毒β属的Sarbecovirus亚属。现在的病毒序列有微弱的时间进化信号,tMRCA平均时间为73 d,95%可信区间(38.9~119.3 d),与BetaCoV/bat/Yunnan/RaTG13/2013病毒不具正性时间进化信号,与bat-SL-CoVZC45和SARS-CoV具有强的正性时间进化关系。病毒在流行期间存在变异,主要是净化选择压力。
结论: 病毒SARS-CoV-2可能出现在2019年11月左右,来源于蝙蝠相关冠状病毒。结果将有助于研究病毒SARS-CoV-2的溯源、进化,对疾病进行正确防控具有指导意义。
Keywords: SARS-CoV-2; coronavirus; evolution; mutation.
Figures


References
-
- Wu F, Zhao S, Yu B, et al. A new coronavirus associated with human respiratory disease in China. https://www.ncbi.nlm.nih.gov/pubmed/32015508. Nature. 2020 doi: 10.1038/s41586-020-2008-3. [Wu F, Zhao S, Yu B, et al. A new coronavirus associated with human respiratory disease in China[J]. Nature, 2020, 10.1038/ s41586-020-2008-3.] - DOI - PMC - PubMed
-
- Zhou P, Yang XL, Wang XG, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. https://www.ncbi.nlm.nih.gov/pubmed/32015507. Nature. 2020 doi: 10.1038/s41586-020-2012-7. [Zhou P, Yang XL, Wang XG, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin[J]. Nature, 2020, 10.1038/s41586-020-2012-7.] - DOI - PMC - PubMed
-
- Rubin EJ, Baden LR, Morrissey S, et al. Medical Journals and the 2019-nCoV Outbreak. http://cn.bing.com/academic/profile?id=3d2f310d5fd9ca41904de4521d724fcc&.... N Engl J Med. 2020 doi: 10.1056/NEJMe2001329. [Rubin EJ, Baden LR, Morrissey S, et al. Medical Journals and the 2019-nCoV Outbreak[J]. N Engl J Med, 2020, 10.1056/NEJMe2001329.] - DOI - PubMed
-
- Guarner J. Three emerging coronaviruses in two decades. https://academic.oup.com/ajcp/article/153/4/420/5735509. Am J Clin Pathol. 2020;aqaa029 [Guarner J. Three emerging coronaviruses in two decades[J]. Am J Clin Pathol, 2020, aqaa029.] - PMC - PubMed
-
- Stein RA. The 2019 Coronavirus: Learning Curves, Lessons, and the Weakest Link. https://www.ncbi.nlm.nih.gov/pubmed/32052918. Int J Clin Pract. 2020:e13488. [Stein RA. The 2019 Coronavirus: Learning Curves, Lessons, and the Weakest Link[J]. Int J Clin Pract, 2020, e13488.] - PMC - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous
