Disruption of ATRX-RNA interactions uncovers roles in ATRX localization and PRC2 function
- PMID: 32376827
- PMCID: PMC7203109
- DOI: 10.1038/s41467-020-15902-9
Disruption of ATRX-RNA interactions uncovers roles in ATRX localization and PRC2 function
Abstract
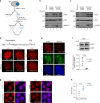
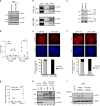
Heterochromatin in the eukaryotic genome is rigorously controlled by the concerted action of protein factors and RNAs. Here, we investigate the RNA binding function of ATRX, a chromatin remodeler with roles in silencing of repetitive regions of the genome and in recruitment of the polycomb repressive complex 2 (PRC2). We identify ATRX RNA binding regions (RBRs) and discover that the major ATRX RBR lies within the N-terminal region of the protein, distinct from its PHD and helicase domains. Deletion of this ATRX RBR (ATRXΔRBR) compromises ATRX interactions with RNAs in vitro and in vivo and alters its chromatin binding properties. Genome-wide studies reveal that loss of RNA interactions results in a redistribution of ATRX on chromatin. Finally, our studies identify a role for ATRX-RNA interactions in regulating PRC2 localization to a subset of polycomb target genes.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
ATRX histone binding and helicase activities have distinct roles in neuronal differentiation.Nucleic Acids Res. 2022 Sep 9;50(16):9162-9174. doi: 10.1093/nar/gkac683. Nucleic Acids Res. 2022. PMID: 35998910 Free PMC article.
-
PML protein organizes heterochromatin domains where it regulates histone H3.3 deposition by ATRX/DAXX.Genome Res. 2017 Jun;27(6):913-921. doi: 10.1101/gr.215830.116. Epub 2017 Mar 24. Genome Res. 2017. PMID: 28341773 Free PMC article.
-
Global changes in chromatin accessibility and transcription following ATRX inactivation in human cancer cells.FEBS Lett. 2020 Jan;594(1):67-78. doi: 10.1002/1873-3468.13549. Epub 2019 Aug 8. FEBS Lett. 2020. PMID: 31329278
-
The recruitment of chromatin modifiers by long noncoding RNAs: lessons from PRC2.RNA. 2015 Dec;21(12):2007-22. doi: 10.1261/rna.053918.115. RNA. 2015. PMID: 26574518 Free PMC article. Review.
-
ATRX and DAXX: Mechanisms and Mutations.Cold Spring Harb Perspect Med. 2017 Mar 1;7(3):a026567. doi: 10.1101/cshperspect.a026567. Cold Spring Harb Perspect Med. 2017. PMID: 28062559 Free PMC article. Review.
Cited by
-
Epigenetic mechanisms regulate sex-specific bias in disease manifestations.J Mol Med (Berl). 2022 Aug;100(8):1111-1123. doi: 10.1007/s00109-022-02227-x. Epub 2022 Jun 29. J Mol Med (Berl). 2022. PMID: 35764820 Free PMC article. Review.
-
Molecular profiling of a real-world breast cancer cohort with genetically inferred ancestries reveals actionable tumor biology differences between European ancestry and African ancestry patient populations.Breast Cancer Res. 2023 May 25;25(1):58. doi: 10.1186/s13058-023-01627-2. Breast Cancer Res. 2023. PMID: 37231433 Free PMC article.
-
Phase separated condensates of ATRX regulate neural progenitor identity.Nat Commun. 2025 Jul 14;16(1):6489. doi: 10.1038/s41467-025-61881-0. Nat Commun. 2025. PMID: 40659667 Free PMC article.
-
Pancreatic Neuroendocrine Tumors: Signaling Pathways and Epigenetic Regulation.Int J Mol Sci. 2024 Jan 22;25(2):1331. doi: 10.3390/ijms25021331. Int J Mol Sci. 2024. PMID: 38279330 Free PMC article. Review.
-
An evolving landscape of PRC2-RNA interactions in chromatin regulation.Nat Rev Mol Cell Biol. 2025 Aug;26(8):631-642. doi: 10.1038/s41580-025-00850-3. Epub 2025 Apr 30. Nat Rev Mol Cell Biol. 2025. PMID: 40307460 Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

