The phosphorelay BarA/SirA activates the non-cognate regulator RcsB in Salmonella enterica
- PMID: 32392214
- PMCID: PMC7241856
- DOI: 10.1371/journal.pgen.1008722
The phosphorelay BarA/SirA activates the non-cognate regulator RcsB in Salmonella enterica
Abstract
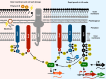
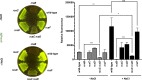
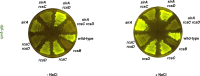
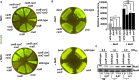
To survive an environmental stress, organisms must detect the stress and mount an appropriate response. One way that bacteria do so is by phosphorelay systems that respond to a stress by activating a regulator that modifies gene expression. To ensure an appropriate response, a given regulator is typically activated solely by its cognate phosphorelay protein(s). However, we now report that the regulator RcsB is activated by both cognate and non-cognate phosphorelay proteins, depending on the condition experienced by the bacterium Salmonella enterica serovar Typhimurium. The RcsC and RcsD proteins form a phosphorelay that activates their cognate regulator RcsB in response to outer membrane stress and cell wall perturbations, conditions Salmonella experiences during infection. Surprisingly, the non-cognate phosphorelay protein BarA activates RcsB during logarithmic growth in Luria-Bertani medium in three ways. That is, BarA's cognate regulator SirA promotes transcription of the rcsDB operon; the SirA-dependent regulatory RNAs CsrB and CsrC further increase RcsB-activated gene transcription; and BarA activates RcsB independently of the RcsC, RcsD, and SirA proteins. Activation of a regulator by multiple sensors broadens the spectrum of environments in which a set of genes is expressed without evolving binding sites for different regulators at each of these genes.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




Similar articles
-
The Rcs phosphorelay system is specific to enteric pathogens/commensals and activates ydeI, a gene important for persistent Salmonella infection of mice.Mol Microbiol. 2006 Nov;62(3):883-94. doi: 10.1111/j.1365-2958.2006.05420.x. Epub 2006 Sep 29. Mol Microbiol. 2006. PMID: 17010160 Free PMC article.
-
Identification of a new promoter for the response regulator rcsB expression in Salmonella enterica serovar Typhimurium.FEMS Microbiol Lett. 2009 Nov;300(2):165-73. doi: 10.1111/j.1574-6968.2009.01771.x. Epub 2009 Aug 28. FEMS Microbiol Lett. 2009. PMID: 19780840
-
Repression of the RcsC-YojN-RcsB phosphorelay by the IgaA protein is a requisite for Salmonella virulence.Mol Microbiol. 2004 Sep;53(5):1437-49. doi: 10.1111/j.1365-2958.2004.04213.x. Mol Microbiol. 2004. PMID: 15387821
-
The Complex Rcs Regulatory Cascade.Annu Rev Microbiol. 2018 Sep 8;72:111-139. doi: 10.1146/annurev-micro-090817-062640. Epub 2018 Jun 13. Annu Rev Microbiol. 2018. PMID: 29897834 Review.
-
The Rcs phosphorelay: more than just a two-component pathway.Future Microbiol. 2010 Aug;5(8):1173-84. doi: 10.2217/fmb.10.83. Future Microbiol. 2010. PMID: 20722597 Review.
Cited by
-
The Rsm (Csr) post-transcriptional regulatory pathway coordinately controls multiple CRISPR-Cas immune systems.Nucleic Acids Res. 2021 Sep 20;49(16):9508-9525. doi: 10.1093/nar/gkab704. Nucleic Acids Res. 2021. PMID: 34403463 Free PMC article.
-
The Rcs stress response inversely controls surface and CRISPR-Cas adaptive immunity to discriminate plasmids and phages.Nat Microbiol. 2021 Feb;6(2):162-172. doi: 10.1038/s41564-020-00822-7. Epub 2021 Jan 4. Nat Microbiol. 2021. PMID: 33398095
-
RcsF-independent mechanisms of signaling within the Rcs phosphorelay.PLoS Genet. 2024 Dec 26;20(12):e1011408. doi: 10.1371/journal.pgen.1011408. eCollection 2024 Dec. PLoS Genet. 2024. PMID: 39724052 Free PMC article.
-
Structural and functional insights underlying recognition of histidine phosphotransfer protein in fungal phosphorelay systems.Commun Biol. 2024 Jul 4;7(1):814. doi: 10.1038/s42003-024-06459-0. Commun Biol. 2024. PMID: 38965424 Free PMC article.
-
HilE is required for synergistic activation of SPI-1 gene expression in Salmonella enterica serovar Typhimurium.BMC Microbiol. 2021 Feb 16;21(1):49. doi: 10.1186/s12866-021-02110-8. BMC Microbiol. 2021. PMID: 33593291 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

