Housefly (Musca domestica L.) associated microbiota across different life stages
- PMID: 32398740
- PMCID: PMC7217826
- DOI: 10.1038/s41598-020-64704-y
Housefly (Musca domestica L.) associated microbiota across different life stages
Abstract
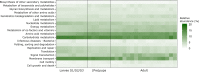
The housefly (Musca domestica L.) lives in close association with its microbiota and its symbionts are suggested to have pivotal roles in processes such as metabolism and immune response, but it is unclear how the profound physiological changes during ontogeny affect the housefly's associated microbiota and their metabolic capabilities. The present study applies 16S rRNA gene amplicon sequencing to investigate the development of the host-associated microbiota during ontogeny. The potential for microbiota transfer between developmental stages, and the metabolic potential of these microbiota were evaluated. Representatives of Firmicutes were observed as early colonisers during the larval stages, followed by colonisation by organisms affiliating with Proteobacteria and Bacteroidetes as the flies matured into adults. Microbiota observed across all the developmental stages included Lactococcus, Lactobacillus and Enterococcus, while Weissella and Chishuiella were associated with newly hatched larvae and adults, respectively. Predictive metabolic profiling of the identified microorganisms further suggested that the microbiota and their functional profile mature alongside their host and putative host-microbe relationships are established at different stages of development. The predicted metabolic capability of the microbiota developed from primarily simple processes including carbohydrate and nucleotide metabolisms, to more complex metabolic pathways including amino acid metabolisms and processes related to signal transduction.
Conflict of interest statement
The authors declare no competing interests.
Figures




References
-
- Engel P, Moran NA. The gut microbiota of insects - diversity in structure and function. FEMS Microbiol. Rev. 2013;37:699–735. - PubMed
-
- Dillon RJ, Dillon VM. The gut bacteria of insects: nonpathogenic interactions. Annu. Rev. Entomol. 2004;49:71–92. - PubMed
-
- Feldhaar H. Bacterial symbionts as mediators of ecologically important traits of insect hosts. Ecol. Entomol. 2011;36:533–543.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

