GGPS1 Mutations Cause Muscular Dystrophy/Hearing Loss/Ovarian Insufficiency Syndrome
- PMID: 32403198
- PMCID: PMC7496979
- DOI: 10.1002/ana.25772
GGPS1 Mutations Cause Muscular Dystrophy/Hearing Loss/Ovarian Insufficiency Syndrome
Abstract
Objective: A hitherto undescribed phenotype of early onset muscular dystrophy associated with sensorineural hearing loss and primary ovarian insufficiency was initially identified in 2 siblings and in subsequent patients with a similar constellation of findings. The goal of this study was to understand the genetic and molecular etiology of this condition.
Methods: We applied whole exome sequencing (WES) superimposed on shared haplotype regions to identify the initial biallelic variants in GGPS1 followed by GGPS1 Sanger sequencing or WES in 5 additional families with the same phenotype. Molecular modeling, biochemical analysis, laser membrane injury assay, and the generation of a Y259C knock-in mouse were done.
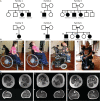
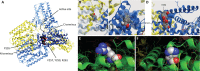
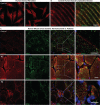
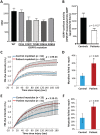
Results: A total of 11 patients in 6 families carrying 5 different biallelic pathogenic variants in specific domains of GGPS1 were identified. GGPS1 encodes geranylgeranyl diphosphate synthase in the mevalonate/isoprenoid pathway, which catalyzes the synthesis of geranylgeranyl pyrophosphate, the lipid precursor of geranylgeranylated proteins including small guanosine triphosphatases. In addition to proximal weakness, all but one patient presented with congenital sensorineural hearing loss, and all postpubertal females had primary ovarian insufficiency. Muscle histology was dystrophic, with ultrastructural evidence of autophagic material and large mitochondria in the most severe cases. There was delayed membrane healing after laser injury in patient-derived myogenic cells, and a knock-in mouse of one of the mutations (Y259C) resulted in prenatal lethality.
Interpretation: The identification of specific GGPS1 mutations defines the cause of a unique form of muscular dystrophy with hearing loss and ovarian insufficiency and points to a novel pathway for this clinical constellation. ANN NEUROL 2020;88:332-347.
© 2020 The Authors. Annals of Neurology published by Wiley Periodicals, Inc. on behalf of American Neurological Association.
Conflict of interest statement
T.V. reports personal fees from DZHK, Antisense Therapeutics, Biophytis, Capricor, Italfarmaco, Santhera, Servier, Sarepta, Solid Biosciences, Dynacure, DiNAQOR, and Catabasis outside the submitted work.
Figures


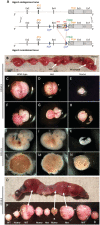
Similar articles
-
GGPS1-associated muscular dystrophy with and without hearing loss.Ann Clin Transl Neurol. 2022 Sep;9(9):1465-1474. doi: 10.1002/acn3.51633. Epub 2022 Jul 23. Ann Clin Transl Neurol. 2022. PMID: 35869884 Free PMC article.
-
Expanding the phenotypic and genotypic spectrum of GGPS1 related congenital muscular dystrophy.Am J Med Genet A. 2024 Apr;194(4):e63498. doi: 10.1002/ajmg.a.63498. Epub 2023 Dec 21. Am J Med Genet A. 2024. PMID: 38129970
-
Genomic sequencing highlights the diverse molecular causes of Perrault syndrome: a peroxisomal disorder (PEX6), metabolic disorders (CLPP, GGPS1), and mtDNA maintenance/translation disorders (LARS2, TFAM).Hum Genet. 2020 Oct;139(10):1325-1343. doi: 10.1007/s00439-020-02176-w. Epub 2020 May 12. Hum Genet. 2020. PMID: 32399598
-
Functional Characterization of a GGPPS Variant Identified in Atypical Femoral Fracture Patients and Delineation of the Role of GGPPS in Bone-Relevant Cell Types.J Bone Miner Res. 2018 Dec;33(12):2091-2098. doi: 10.1002/jbmr.3580. Epub 2018 Sep 24. J Bone Miner Res. 2018. PMID: 30184270
-
Recessive LOXHD1 variants cause a prelingual down-sloping hearing loss: genotype-phenotype correlation and three additional children with novel variants.Int J Pediatr Otorhinolaryngol. 2021 Jun;145:110715. doi: 10.1016/j.ijporl.2021.110715. Epub 2021 Apr 20. Int J Pediatr Otorhinolaryngol. 2021. PMID: 33892339 Review.
Cited by
-
Two Novel Pathogenic Variants Confirm RMND1 Causative Role in Perrault Syndrome with Renal Involvement.Genes (Basel). 2020 Sep 8;11(9):1060. doi: 10.3390/genes11091060. Genes (Basel). 2020. PMID: 32911714 Free PMC article.
-
Whole-Genome Scanning for Selection Signatures Reveals Candidate Genes Associated with Growth and Tail Length in Sheep.Animals (Basel). 2024 Feb 22;14(5):687. doi: 10.3390/ani14050687. Animals (Basel). 2024. PMID: 38473071 Free PMC article.
-
HMGCS1 variants cause rigid spine syndrome amenable to mevalonic acid treatment in an animal model.Brain. 2025 May 13;148(5):1707-1722. doi: 10.1093/brain/awae371. Brain. 2025. PMID: 39531736
-
New insights into Perrault syndrome, a clinically and genetically heterogeneous disorder.Hum Genet. 2022 Apr;141(3-4):805-819. doi: 10.1007/s00439-021-02319-7. Epub 2021 Aug 2. Hum Genet. 2022. PMID: 34338890 Free PMC article. Review.
-
Geranylgeranyl diphosphate synthase: Role in human health, disease and potential therapeutic target.Clin Transl Med. 2023 Jan;13(1):e1167. doi: 10.1002/ctm2.1167. Clin Transl Med. 2023. PMID: 36650113 Free PMC article. Review.
References
-
- Mercuri E, Bonnemann CG, Muntoni F. Muscular dystrophies. Lancet 2019;394:2025–2038. - PubMed
-
- Tawil R. Facioscapulohumeral muscular dystrophy. Handb Clin Neurol 2018;148:541–548. - PubMed
-
- Demain LA, Urquhart JE, O'Sullivan J, et al. Expanding the genotypic spectrum of Perrault syndrome. Clin Genet 2017;91:302–312. - PubMed
-
- Goldstein JL, Brown MS. Regulation of the mevalonate pathway. Nature 1990;343:425–430. - PubMed
-
- Xu N, Shen N, Wang X, et al. Protein prenylation and human diseases: a balance of protein farnesylation and geranylgeranylation. Sci China Life Sci 2015;58:328–335. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

