Mass spectrometry for the identification and analysis of highly complex glycosylation of therapeutic or pathogenic proteins
- PMID: 32406805
- PMCID: PMC7372739
- DOI: 10.1080/14789450.2020.1769479
Mass spectrometry for the identification and analysis of highly complex glycosylation of therapeutic or pathogenic proteins
Abstract
Introduction: Protein glycosylation influences characteristics such as folding, stability, protein interactions, and solubility. Therefore, glycan moieties of therapeutic proteins and proteins that are likely associated with disease pathogenesis should be analyzed in-depth, including glycan heterogeneity and modification sites. Recent advances in analytical methods and instrumentation have enabled comprehensive characterization of highly complex glycosylated proteins.
Area covered: The following aspects should be considered when analyzing glycosylated proteins: sample preparation, chromatographic separation, mass spectrometry (MS) and fragmentation methods, and bioinformatics, such as software solutions for data analyses. Notably, analysis of glycoproteins with heavily sialylated glycans or multiple glycosylation sites requires special considerations. Here, we discuss recent methodological advances in MS that provide detailed characterization of heterogeneous glycoproteins.
Expert opinion: As characterization of complex glycosylated proteins is still analytically challenging, the function or pathophysiological significance of these proteins is not fully understood. To reproducibly produce desired forms of therapeutic glycoproteins or to fully elucidate disease-specific patterns of protein glycosylation, a highly reproducible and robust analytical platform(s) should be established. In addition to advances in MS instrumentation, optimization of analytical and bioinformatics methods and utilization of glycoprotein/glycopeptide standards is desirable. Ultimately, we envision that an automated high-throughput MS analysis will provide additional power to clinical studies and precision medicine.
Keywords: N-glycosylation; O-glycosylation; Fc fusions protein therapeutics; Mucin 1 (MUC-1); erythropoietin; immunoglobulin glycosylation; intravenous immunoglobulin (IVIG); virus glycoconjugates.
Conflict of interest statement
Disclosure of interests
The authors have no relevant affiliation or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript. In the interest of full disclosure, we report that M. B. Renfrow and J. Novak are co-founders of Reliant Glycosciences, LLC and co‑inventors on the US patent application 14/318,082 (assigned to UAB Research Foundation that distributes royalties to the inventors).
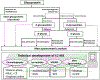
Figures




Similar articles
-
Mass Spectrometry-Based Chemical and Enzymatic Methods for Global Analysis of Protein Glycosylation.Acc Chem Res. 2018 Aug 21;51(8):1796-1806. doi: 10.1021/acs.accounts.8b00200. Epub 2018 Jul 16. Acc Chem Res. 2018. PMID: 30011186 Free PMC article. Review.
-
Carbohydrates on Proteins: Site-Specific Glycosylation Analysis by Mass Spectrometry.Annu Rev Anal Chem (Palo Alto Calif). 2015;8:463-83. doi: 10.1146/annurev-anchem-071114-040240. Epub 2015 Jun 11. Annu Rev Anal Chem (Palo Alto Calif). 2015. PMID: 26070719 Review.
-
Systems analysis of singly and multiply O-glycosylated peptides in the human serum glycoproteome via EThcD and HCD mass spectrometry.J Proteomics. 2018 Jan 6;170:14-27. doi: 10.1016/j.jprot.2017.09.014. Epub 2017 Sep 29. J Proteomics. 2018. PMID: 28970103
-
Recent advances in the MS analysis of glycoproteins: Theoretical considerations.Electrophoresis. 2011 Jan;32(1):3-13. doi: 10.1002/elps.201000393. Epub 2010 Nov 30. Electrophoresis. 2011. PMID: 21171109 Free PMC article. Review.
-
Intact glycopeptide characterization using mass spectrometry.Expert Rev Proteomics. 2016 May;13(5):513-22. doi: 10.1586/14789450.2016.1172965. Expert Rev Proteomics. 2016. PMID: 27140194 Free PMC article. Review.
Cited by
-
The role of metagenomic next-generation sequencing in diagnosing and managing post-kidney transplantation infections.Front Cell Infect Microbiol. 2025 Jan 7;14:1473068. doi: 10.3389/fcimb.2024.1473068. eCollection 2024. Front Cell Infect Microbiol. 2025. PMID: 39839264 Free PMC article. Review.
-
Mannose-specific plant and microbial lectins as antiviral agents: A review.Glycoconj J. 2024 Feb;41(1):1-33. doi: 10.1007/s10719-023-10142-7. Epub 2024 Jan 20. Glycoconj J. 2024. PMID: 38244136 Review.
-
High sensitivity glycomics in biomedicine.Mass Spectrom Rev. 2022 Nov;41(6):1014-1039. doi: 10.1002/mas.21730. Epub 2021 Sep 7. Mass Spectrom Rev. 2022. PMID: 34494287 Free PMC article. Review.
-
Immunoglobulin A Glycosylation and Its Role in Disease.Exp Suppl. 2021;112:433-477. doi: 10.1007/978-3-030-76912-3_14. Exp Suppl. 2021. PMID: 34687019
-
O-glycosylation of IgA1 and the pathogenesis of an autoimmune disease IgA nephropathy.Glycobiology. 2024 Sep 30;34(11):cwae060. doi: 10.1093/glycob/cwae060. Glycobiology. 2024. PMID: 39095059 Free PMC article. Review.
References
-
-
Reily C, Stewart TJ, Renfrow MB, Novak J. Glycosylation in health and disease. Nat Rev Nephrol. 2019;15(6):346–66. doi: 10.1038/s41581-019-0129-4.
** This review provides fundamental information of glycosylation pathway and shows how glycosylation patterns are altered in multiple human diseases.
-
-
-
O’Flaherty R, Trbojevic-Akmacic I, Greville G, Rudd PM, Lauc G. The sweet spot for biologics: recent advances in characterization of biotherapeutic glycoproteins. Expert Rev Proteomics. 2018;15(1):13–29. doi: 10.1080/14789450.2018.1404907.
* This review highlights recent advances in analytical technologies for characterization of biotherapeutic glycoproteins.
-
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
