Escherichia coli YcaQ is a DNA glycosylase that unhooks DNA interstrand crosslinks
- PMID: 32409837
- PMCID: PMC7367128
- DOI: 10.1093/nar/gkaa346
Escherichia coli YcaQ is a DNA glycosylase that unhooks DNA interstrand crosslinks
Abstract
Interstrand DNA crosslinks (ICLs) are a toxic form of DNA damage that block DNA replication and transcription by tethering the opposing strands of DNA. ICL repair requires unhooking of the tethered strands by either nuclease incision of the DNA backbone or glycosylase cleavage of the crosslinked nucleotide. In bacteria, glycosylase-mediated ICL unhooking was described in Streptomyces as a means of self-resistance to the genotoxic natural product azinomycin B. The mechanistic details and general utility of glycosylase-mediated ICL repair in other bacteria are unknown. Here, we identify the uncharacterized Escherichia coli protein YcaQ as an ICL repair glycosylase that protects cells against the toxicity of crosslinking agents. YcaQ unhooks both sides of symmetric and asymmetric ICLs in vitro, and loss or overexpression of ycaQ sensitizes E. coli to the nitrogen mustard mechlorethamine. Comparison of YcaQ and UvrA-mediated ICL resistance mechanisms establishes base excision as an alternate ICL repair pathway in bacteria.
© The Author(s) 2020. Published by Oxford University Press on behalf of Nucleic Acids Research.
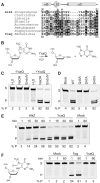
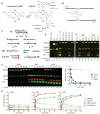
Figures




References
-
- Scharer O.D. DNA interstrand crosslinks: natural and drug-induced DNA adducts that induce unique cellular responses. ChemBioChem. 2005; 6:27–32. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

