High Levels of Chromosomal Copy Number Alterations and TP53 Mutations Correlate with Poor Outcome in Younger Breast Cancer Patients
- PMID: 32416097
- PMCID: PMC7397461
- DOI: 10.1016/j.ajpath.2020.04.015
High Levels of Chromosomal Copy Number Alterations and TP53 Mutations Correlate with Poor Outcome in Younger Breast Cancer Patients
Abstract
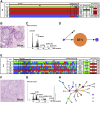
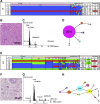
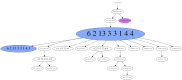
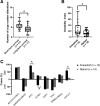
Prognosis in young patients with breast cancer is generally poor, yet considerable differences in clinical outcomes between individual patients exist. To understand the genetic basis of the disparate clinical courses, tumors were collected from 34 younger women, 17 with good and 17 with poor outcomes, as determined by disease-specific survival during a follow-up period of 17 years. The clinicopathologic parameters of the tumors were complemented with DNA image cytometry profiles, enumeration of copy numbers of eight breast cancer genes by multicolor fluorescence in situ hybridization, and targeted sequence analysis of 563 cancer genes. Both groups included diploid and aneuploid tumors. The degree of intratumor heterogeneity was significantly higher in aneuploid versus diploid cases, and so were gains of the oncogenes MYC and ZNF217. Significantly more copy number alterations were observed in the group with poor outcome. Almost all tumors in the group with long survival were classified as luminal A, whereas triple-negative tumors predominantly occurred in the short survival group. Mutations in PIK3CA were more common in the group with good outcome, whereas TP53 mutations were more frequent in patients with poor outcomes. This study shows that TP53 mutations and the extent of genomic imbalances are associated with poor outcome in younger breast cancer patients and thus emphasize the central role of genomic instability vis-a-vis tumor aggressiveness.
Copyright © 2020 American Society for Investigative Pathology. Published by Elsevier Inc. All rights reserved.
Figures

Similar articles
-
Single Cell Genetic Profiling of Tumors of Breast Cancer Patients Aged 50 Years and Older Reveals Enormous Intratumor Heterogeneity Independent of Individual Prognosis.Cancers (Basel). 2021 Jul 5;13(13):3366. doi: 10.3390/cancers13133366. Cancers (Basel). 2021. PMID: 34282768 Free PMC article.
-
Aneuploidy, TP53 mutation, and amplification of MYC correlate with increased intratumor heterogeneity and poor prognosis of breast cancer patients.Genes Chromosomes Cancer. 2018 Apr;57(4):165-175. doi: 10.1002/gcc.22515. Epub 2018 Jan 9. Genes Chromosomes Cancer. 2018. PMID: 29181861 Free PMC article.
-
Inter-tumor genomic heterogeneity of breast cancers: comprehensive genomic profile of primary early breast cancers and relapses.Breast Cancer Res. 2020 Oct 15;22(1):107. doi: 10.1186/s13058-020-01345-z. Breast Cancer Res. 2020. PMID: 33059724 Free PMC article.
-
p53 in breast cancer subtypes and new insights into response to chemotherapy.Breast. 2013 Aug;22 Suppl 2:S27-9. doi: 10.1016/j.breast.2013.07.005. Breast. 2013. PMID: 24074787 Review.
-
The Unique Biology behind the Early Onset of Breast Cancer.Genes (Basel). 2021 Mar 5;12(3):372. doi: 10.3390/genes12030372. Genes (Basel). 2021. PMID: 33807872 Free PMC article. Review.
Cited by
-
Single Cell Genetic Profiling of Tumors of Breast Cancer Patients Aged 50 Years and Older Reveals Enormous Intratumor Heterogeneity Independent of Individual Prognosis.Cancers (Basel). 2021 Jul 5;13(13):3366. doi: 10.3390/cancers13133366. Cancers (Basel). 2021. PMID: 34282768 Free PMC article.
-
Genomic characterization of the HER2-enriched intrinsic molecular subtype in primary ER-positive HER2-negative breast cancer.Nat Commun. 2025 Mar 5;16(1):2208. doi: 10.1038/s41467-025-57419-z. Nat Commun. 2025. PMID: 40044693 Free PMC article.
-
Evaluating the prognostic potential of telomerase signature in breast cancer through advanced machine learning model.Front Immunol. 2024 Nov 28;15:1462953. doi: 10.3389/fimmu.2024.1462953. eCollection 2024. Front Immunol. 2024. PMID: 39669558 Free PMC article.
-
Copy number alteration is an independent prognostic biomarker in triple-negative breast cancer patients.Breast Cancer. 2023 Jul;30(4):584-595. doi: 10.1007/s12282-023-01449-2. Epub 2023 Mar 17. Breast Cancer. 2023. PMID: 36930419
-
Prognostic Value of the TP53 Mutation Location in Metastatic Breast Cancer as Detected by Next-Generation Sequencing.Cancer Manag Res. 2021 Apr 15;13:3303-3316. doi: 10.2147/CMAR.S298729. eCollection 2021. Cancer Manag Res. 2021. PMID: 33889023 Free PMC article.
References
-
- Bray F., Ferlay J., Soerjomataram I., Siegel R.L., Torre L.A., Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424. - PubMed
-
- Adami H.O., Malker B., Holmberg L., Persson I., Stone B. The relation between survival and age at diagnosis in breast cancer. N Engl J Med. 1986;315:559–563. - PubMed
-
- Nixon A.J., Neuberg D., Hayes D.F., Gelman R., Connolly J.L., Schnitt S., Abner A., Recht A., Vicini F., Harris J.R. Relationship of patient age to pathologic features of the tumor and prognosis for patients with stage I or II breast cancer. J Clin Oncol. 1994;12:888–894. - PubMed
-
- Sariego J. Breast cancer in the young patient. Am Surg. 2010;76:1397–1400. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

