Polymorphic centromere locations in the pathogenic yeast Candida parapsilosis
- PMID: 32424070
- PMCID: PMC7263194
- DOI: 10.1101/gr.257816.119
Polymorphic centromere locations in the pathogenic yeast Candida parapsilosis
Abstract
Centromeres pose an evolutionary paradox: strongly conserved in function but rapidly changing in sequence and structure. However, in the absence of damage, centromere locations are usually conserved within a species. We report here that isolates of the pathogenic yeast species Candida parapsilosis show within-species polymorphism for the location of centromeres on two of its eight chromosomes. Its old centromeres have an inverted-repeat (IR) structure, whereas its new centromeres have no obvious structural features but are located within 30 kb of the old site. Centromeres can therefore move naturally from one chromosomal site to another, apparently spontaneously and in the absence of any significant changes in DNA sequence. Our observations are consistent with a model in which all centromeres are genetically determined, such as by the presence of short or long IRs or by the ability to form cruciforms. We also find that centromeres have been hotspots for genomic rearrangements in the C. parapsilosis clade.
© 2020 Ola et al.; Published by Cold Spring Harbor Laboratory Press.
Figures


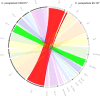



References
Publication types
MeSH terms
Supplementary concepts
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
