Switching the Switch: Ligand Induced Disulfide Formation in HDAC8
- PMID: 32428298
- PMCID: PMC7692948
- DOI: 10.1002/chem.202001712
Switching the Switch: Ligand Induced Disulfide Formation in HDAC8
Abstract
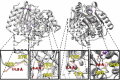
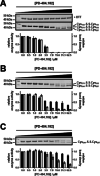
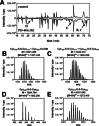
Human histone deacetylase 8 is a well-recognized target for T-cell lymphoma and particularly childhood neuroblastoma. PD-404,182 was shown to be a selective covalent inhibitor of HDAC8 that forms mixed disulfides with several cysteine residues and is also able to transform thiol groups to thiocyanates. Moreover, HDAC8 was shown to be regulated by a redox switch based on the reversible formation of a disulfide bond between cysteines Cys102 and Cys153 . This study on the distinct effects of PD-404,182 on HDAC8 reveals that this compound induces the dose-dependent formation of intramolecular disulfide bridges. Therefore, the inhibition mechanism of HDAC8 by PD-404,182 involves both, covalent modification of thiols as well as ligand mediated disulfide formation. Moreover, this study provides a deep molecular insight into the regulation mechanism of HDAC8 involving several cysteines with graduated capability to form reversible disulfide bridges.
Keywords: HDAC8; covalent inhibitors; cysteine; redox switch; sulfenamides.
© 2020 The Authors. Published by Wiley-VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
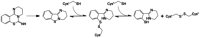

Similar articles
-
3-Chloro-5-Substituted-1,2,4-Thiadiazoles (TDZs) as Selective and Efficient Protein Thiol Modifiers.Chembiochem. 2022 Nov 4;23(21):e202200417. doi: 10.1002/cbic.202200417. Epub 2022 Sep 27. Chembiochem. 2022. PMID: 36066474 Free PMC article.
-
Covalent inhibition of histone deacetylase 8 by 3,4-dihydro-2H-pyrimido[1,2-c][1,3]benzothiazin-6-imine.Biochim Biophys Acta Gen Subj. 2019 Mar;1863(3):577-585. doi: 10.1016/j.bbagen.2019.01.001. Epub 2019 Jan 3. Biochim Biophys Acta Gen Subj. 2019. PMID: 30611847
-
The enzyme activity of histone deacetylase 8 is modulated by a redox-switch.Redox Biol. 2019 Jan;20:60-67. doi: 10.1016/j.redox.2018.09.013. Epub 2018 Sep 27. Redox Biol. 2019. PMID: 30292946 Free PMC article.
-
Targeting of HDAC8 and investigational inhibitors in neuroblastoma.Expert Opin Investig Drugs. 2009 Nov;18(11):1605-17. doi: 10.1517/14728220903241658. Expert Opin Investig Drugs. 2009. PMID: 19780707 Review.
-
Histone deacetylase 8 (HDAC8) and its inhibitors with selectivity to other isoforms: An overview.Eur J Med Chem. 2019 Feb 15;164:214-240. doi: 10.1016/j.ejmech.2018.12.039. Epub 2018 Dec 19. Eur J Med Chem. 2019. PMID: 30594678 Review.
Cited by
-
Covalent Inhibition by a Natural Product-Inspired Latent Electrophile.J Am Chem Soc. 2023 May 24;145(20):11097-11109. doi: 10.1021/jacs.3c00598. Epub 2023 May 15. J Am Chem Soc. 2023. PMID: 37183434 Free PMC article.
-
Epigenetic alterations and advancement of lymphoma treatment.Ann Hematol. 2024 May;103(5):1435-1454. doi: 10.1007/s00277-023-05395-z. Epub 2023 Aug 15. Ann Hematol. 2024. PMID: 37581713 Review.
-
Rapid Determination of Kinetic Constants for Slow-Binding Inhibitors and Inactivators of Human Histone Deacetylase 8.Int J Mol Sci. 2024 May 21;25(11):5593. doi: 10.3390/ijms25115593. Int J Mol Sci. 2024. PMID: 38891780 Free PMC article.
-
3-Chloro-5-Substituted-1,2,4-Thiadiazoles (TDZs) as Selective and Efficient Protein Thiol Modifiers.Chembiochem. 2022 Nov 4;23(21):e202200417. doi: 10.1002/cbic.202200417. Epub 2022 Sep 27. Chembiochem. 2022. PMID: 36066474 Free PMC article.
-
Electrophilic MiniFrags Revealed Unprecedented Binding Sites for Covalent HDAC8 Inhibitors.J Med Chem. 2024 Jan 11;67(1):572-585. doi: 10.1021/acs.jmedchem.3c01779. Epub 2023 Dec 19. J Med Chem. 2024. PMID: 38113354 Free PMC article.
References
-
- Deardorff M. A., Bando M., Nakato R., Watrin E., Itoh T., Minamino M., Saitoh K., Komata M., Katou Y., Clark D., Cole K. E., de Baere E., Decroos C., Di Donato N., Ernst S., Francey L. J., Gyftodimou Y., Hirashima K., Hullings M., Ishikawa Y., Jaulin C., Kaur M., Kiyono T., Lombardi P. M., Magnaghi-Jaulin L., Mortier G. R., Nozaki N., Petersen M. B., Seimiya H., Siu V. M., Suzuki Y., Takagaki K., Wilde J. J., Willems P. J., Prigent C., Gillessen-Kaesbach G., Christianson D. W., Kaiser F. J., Jackson L. G., Hirota T., Krantz I. D., Shirahige K., Nature 2012, 489, 313–317. - PMC - PubMed
LinkOut - more resources
Full Text Sources

