Automatic spatial estimation of white matter hyperintensities evolution in brain MRI using disease evolution predictor deep neural networks
- PMID: 32428823
- PMCID: PMC7294240
- DOI: 10.1016/j.media.2020.101712
Automatic spatial estimation of white matter hyperintensities evolution in brain MRI using disease evolution predictor deep neural networks
Abstract
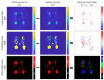
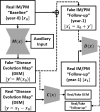
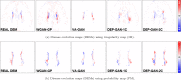
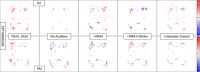
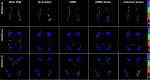
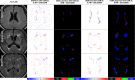
Previous studies have indicated that white matter hyperintensities (WMH), the main radiological feature of small vessel disease, may evolve (i.e., shrink, grow) or stay stable over a period of time. Predicting these changes are challenging because it involves some unknown clinical risk factors that leads to a non-deterministic prediction task. In this study, we propose a deep learning model to predict the evolution of WMH from baseline to follow-up (i.e., 1-year later), namely "Disease Evolution Predictor" (DEP) model, which can be adjusted to become a non-deterministic model. The DEP model receives a baseline image as input and produces a map called "Disease Evolution Map" (DEM), which represents the evolution of WMH from baseline to follow-up. Two DEP models are proposed, namely DEP-UResNet and DEP-GAN, which are representatives of the supervised (i.e., need expert-generated manual labels to generate the output) and unsupervised (i.e., do not require manual labels produced by experts) deep learning algorithms respectively. To simulate the non-deterministic and unknown parameters involved in WMH evolution, we modulate a Gaussian noise array to the DEP model as auxiliary input. This forces the DEP model to imitate a wider spectrum of alternatives in the prediction results. The alternatives of using other types of auxiliary input instead, such as baseline WMH and stroke lesion loads are also proposed and tested. Based on our experiments, the fully supervised machine learning scheme DEP-UResNet regularly performed better than the DEP-GAN which works in principle without using any expert-generated label (i.e., unsupervised). However, a semi-supervised DEP-GAN model, which uses probability maps produced by a supervised segmentation method in the learning process, yielded similar performances to the DEP-UResNet and performed best in the clinical evaluation. Furthermore, an ablation study showed that an auxiliary input, especially the Gaussian noise, improved the performance of DEP models compared to DEP models that lacked the auxiliary input regardless of the model's architecture. To the best of our knowledge, this is the first extensive study on modelling WMH evolution using deep learning algorithms, which deals with the non-deterministic nature of WMH evolution.
Keywords: DEP Generative adversarial network (DEP-GAN); DEP U-Residual Network (DEP-UResNet); Disease evolution predictor (DEP) models; Small vessel disease (SVD); WMH Evolution; White matter hyperintensities (WMH).
Copyright © 2020. Published by Elsevier B.V.
Figures





Similar articles
-
Limited One-time Sampling Irregularity Map (LOTS-IM) for Automatic Unsupervised Assessment of White Matter Hyperintensities and Multiple Sclerosis Lesions in Structural Brain Magnetic Resonance Images.Comput Med Imaging Graph. 2020 Jan;79:101685. doi: 10.1016/j.compmedimag.2019.101685. Epub 2019 Nov 27. Comput Med Imaging Graph. 2020. PMID: 31846826
-
White matter hyperintensity and stroke lesion segmentation and differentiation using convolutional neural networks.Neuroimage Clin. 2017 Dec 20;17:918-934. doi: 10.1016/j.nicl.2017.12.022. eCollection 2018. Neuroimage Clin. 2017. PMID: 29527496 Free PMC article.
-
Segmentation of white matter hyperintensities using convolutional neural networks with global spatial information in routine clinical brain MRI with none or mild vascular pathology.Comput Med Imaging Graph. 2018 Jun;66:28-43. doi: 10.1016/j.compmedimag.2018.02.002. Epub 2018 Feb 17. Comput Med Imaging Graph. 2018. PMID: 29523002
-
Prediction of white matter hyperintensities evolution one-year post-stroke from a single-point brain MRI and stroke lesions information.Sci Rep. 2025 Jan 7;15(1):1208. doi: 10.1038/s41598-024-83128-6. Sci Rep. 2025. PMID: 39774013 Free PMC article.
-
Automatic segmentation of white matter hyperintensities: validation and comparison with state-of-the-art methods on both Multiple Sclerosis and elderly subjects.Neuroimage Clin. 2022;33:102940. doi: 10.1016/j.nicl.2022.102940. Epub 2022 Jan 10. Neuroimage Clin. 2022. PMID: 35051744 Free PMC article.
Cited by
-
Pioneering new paths: the role of generative modelling in neurological disease research.Pflugers Arch. 2025 Apr;477(4):571-589. doi: 10.1007/s00424-024-03016-w. Epub 2024 Oct 8. Pflugers Arch. 2025. PMID: 39377960 Free PMC article. Review.
-
AI in Neurology: Everything, Everywhere, All at Once Part 1: Principles and Practice.Ann Neurol. 2025 Aug;98(2):211-230. doi: 10.1002/ana.27225. Epub 2025 Jun 19. Ann Neurol. 2025. PMID: 40536356 Free PMC article. Review.
-
Individualised prediction of longitudinal change in multimodal brain imaging.Imaging Neurosci (Camb). 2024 Jul 3;2:1-19. doi: 10.1162/imag_a_00215. eCollection 2024 Jul 1. Imaging Neurosci (Camb). 2024. PMID: 40046980 Free PMC article.
-
Small Vessel Disease, a Marker of Brain Health: What the Radiologist Needs to Know.AJNR Am J Neuroradiol. 2022 May;43(5):650-660. doi: 10.3174/ajnr.A7302. Epub 2021 Oct 7. AJNR Am J Neuroradiol. 2022. PMID: 34620594 Free PMC article. Review.
-
Applications of generative adversarial networks in neuroimaging and clinical neuroscience.Neuroimage. 2023 Apr 1;269:119898. doi: 10.1016/j.neuroimage.2023.119898. Epub 2023 Jan 24. Neuroimage. 2023. PMID: 36702211 Free PMC article. Review.
References
-
- Baumgartner C.F., Koch L.M., Tezcan K.C., Ang J.X., Konukoglu E. Proc IEEE Comput Soc Conf Comput Vis Pattern Recognit. 2017. Visual feature attribution using wasserstein gans.
-
- Bigolin Lanfredi R., Schroeder J.D., Vachet C., Tasdizen T. Adversarial Regression Training for Visualizing the Progression of Chronic Obstructive Pulmonary Disease with Chest X-Rays. In: Shen D., editor. Medical Image Computing and Computer Assisted Intervention – MICCAI 2019. MICCAI 2019. vol. 11769. Springer; Cham: 2019. (Lecture Notes in Computer Science). - DOI
-
- Bland J.M., Altman D. Statistical methods for assessing agreement between two methods of clinical measurement. The Lancet. 1986;327(8476):307–310. - PubMed
-
- del C. Valdés Hernández M., Piper R.J., Wang X., Deary I.J., Wardlaw J.M. Towards the automatic computational assessment of enlarged perivascular spaces on brain magnetic resonance images: a systematic review. J. Magn. Reson. Imaging. 2013;38(4):774–785. - PubMed
-
- Chappell F.M., Hernández M.d.C.V., Makin S.D., Shuler K., Sakka E., Dennis M.S., Armitage P.A., Maniega S.M., Wardlaw J.M. Sample size considerations for trials using cerebral white matter hyperintensity progression as an intermediate outcome at 1 year after mild stroke: results of a prospective cohort study. Trials. 2017;18(1):78. - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

