Metabolite Genome-Wide Association Study (mGWAS) and Gene-Metabolite Interaction Network Analysis Reveal Potential Biomarkers for Feed Efficiency in Pigs
- PMID: 32429265
- PMCID: PMC7281523
- DOI: 10.3390/metabo10050201
Metabolite Genome-Wide Association Study (mGWAS) and Gene-Metabolite Interaction Network Analysis Reveal Potential Biomarkers for Feed Efficiency in Pigs
Abstract
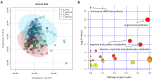
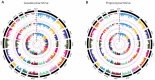
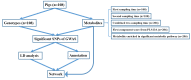
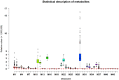
Metabolites represent the ultimate response of biological systems, so metabolomics is considered the link between genotypes and phenotypes. Feed efficiency is one of the most important phenotypes in sustainable pig production and is the main breeding goal trait. We utilized metabolic and genomic datasets from a total of 108 pigs from our own previously published studies that involved 59 Duroc and 49 Landrace pigs with data on feed efficiency (residual feed intake (RFI)), genotype (PorcineSNP80 BeadChip) data, and metabolomic data (45 final metabolite datasets derived from LC-MS system). Utilizing these datasets, our main aim was to identify genetic variants (single-nucleotide polymorphisms (SNPs)) that affect 45 different metabolite concentrations in plasma collected at the start and end of the performance testing of pigs categorized as high or low in their feed efficiency (based on RFI values). Genome-wide significant genetic variants could be then used as potential genetic or biomarkers in breeding programs for feed efficiency. The other objective was to reveal the biochemical mechanisms underlying genetic variation for pigs' feed efficiency. In order to achieve these objectives, we firstly conducted a metabolite genome-wide association study (mGWAS) based on mixed linear models and found 152 genome-wide significant SNPs (p-value < 1.06 × 10-6) in association with 17 metabolites that included 90 significant SNPs annotated to 52 genes. On chromosome one alone, 51 significant SNPs associated with isovalerylcarnitine and propionylcarnitine were found to be in strong linkage disequilibrium (LD). SNPs in strong LD annotated to FBXL4, and CCNC consisted of two haplotype blocks where three SNPs (ALGA0004000, ALGA0004041, and ALGA0004042) were in the intron regions of FBXL4 and CCNC. The interaction network revealed that CCNC and FBXL4 were linked by the hub gene N6AMT1 that was associated with isovalerylcarnitine and propionylcarnitine. Moreover, three metabolites (i.e., isovalerylcarnitine, propionylcarnitine, and pyruvic acid) were clustered in one group based on the low-high RFI pigs. This study performed a comprehensive metabolite-based genome-wide association study (GWAS) analysis for pigs with differences in feed efficiency and provided significant metabolites for which there is significant genetic variation as well as biological interaction networks. The identified metabolite genetic variants, genes, and networks in high versus low feed efficient pigs could be considered as potential genetic or biomarkers for feed efficiency.
Keywords: GWAS; RFI; SNPs; biomarkers; feed efficiency; metabolomics; pathways; pigs.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
Figures



Similar articles
-
Genome-Wide Epistatic Interaction Networks Affecting Feed Efficiency in Duroc and Landrace Pigs.Front Genet. 2020 Feb 28;11:121. doi: 10.3389/fgene.2020.00121. eCollection 2020. Front Genet. 2020. PMID: 32184802 Free PMC article.
-
Genome-wide association and systems genetic analyses of residual feed intake, daily feed consumption, backfat and weight gain in pigs.BMC Genet. 2014 Feb 17;15:27. doi: 10.1186/1471-2156-15-27. BMC Genet. 2014. PMID: 24533460 Free PMC article.
-
Genome-wide association and pathway analysis of feed efficiency in pigs reveal candidate genes and pathways for residual feed intake.Front Genet. 2014 Sep 9;5:307. doi: 10.3389/fgene.2014.00307. eCollection 2014. Front Genet. 2014. PMID: 25250046 Free PMC article.
-
Review: divergent selection for residual feed intake in the growing pig.Animal. 2017 Sep;11(9):1427-1439. doi: 10.1017/S175173111600286X. Epub 2017 Jan 25. Animal. 2017. PMID: 28118862 Free PMC article. Review.
-
Application of GWAS and mGWAS in Livestock and Poultry Breeding.Animals (Basel). 2024 Aug 16;14(16):2382. doi: 10.3390/ani14162382. Animals (Basel). 2024. PMID: 39199916 Free PMC article. Review.
Cited by
-
Assessing Genomic Diversity and Productivity Signatures in Dianzhong Cattle by Whole-Genome Scanning.Front Genet. 2021 Oct 5;12:719215. doi: 10.3389/fgene.2021.719215. eCollection 2021. Front Genet. 2021. PMID: 34675962 Free PMC article.
-
Transcriptomic and metabolomic analysis of autumn leaf color change in Fraxinus angustifolia.PeerJ. 2023 May 12;11:e15319. doi: 10.7717/peerj.15319. eCollection 2023. PeerJ. 2023. PMID: 37197583 Free PMC article.
-
Genetic regulation and variation of fetal plasma metabolome in the context of sex, paternal breeds and variable fetal weight.Open Biol. 2025 Mar;15(3):240285. doi: 10.1098/rsob.240285. Epub 2025 Mar 5. Open Biol. 2025. PMID: 40037532 Free PMC article.
-
Multi-omics insights into key microorganisms and metabolites in Tibetan sheep's high-altitude adaptation.Front Microbiol. 2025 Jun 18;16:1616555. doi: 10.3389/fmicb.2025.1616555. eCollection 2025. Front Microbiol. 2025. PMID: 40606153 Free PMC article.
-
Analysis of merged transcriptomic and genomic datasets to identify genes and pathways underlying residual feed intake in growing pigs.Sci Rep. 2022 Dec 19;12(1):21946. doi: 10.1038/s41598-022-26496-1. Sci Rep. 2022. PMID: 36536008 Free PMC article.
References
-
- Liu H., Chen Y., Ming D., Wang J., Li Z., Ma X., Wang J., Van Milgen J., Wang F. Integrative analysis of indirect calorimetry and metabolomics profiling reveals alterations in energy metabolism between fed and fasted pigs. J. Anim. Sci. Biotechnol. 2018;9:41. doi: 10.1186/s40104-018-0257-x. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

