Metagenomic Diagnosis for a Culture-Negative Sample From a Patient With Severe Pneumonia by Nanopore and Next-Generation Sequencing
- PMID: 32432051
- PMCID: PMC7214676
- DOI: 10.3389/fcimb.2020.00182
Metagenomic Diagnosis for a Culture-Negative Sample From a Patient With Severe Pneumonia by Nanopore and Next-Generation Sequencing
Abstract
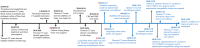
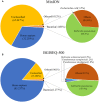
Rapid and accurate etiologic diagnosis accelerates targeted antimicrobial therapy. Metagenomic analysis has played a critical role in pathogen identification. In this study, we leveraged the advantages of both the MinION and BGISEQ-500 platforms to make a bacteriologic diagnosis from a culture-negative lung tissue sample from an immunocompromised patient with severe pneumonia. Real-time nanopore sequencing rapidly identified Klebsiella pneumoniae by an 823 bp specific sequence within 1 min. Genomic analysis further identified blaSHV-12, blaKPC-2, blaTEM-1, blaCTX-M-65, and other resistance genes. The same sample was further sequenced on the BGISEQ-500 platform, which presented consistent results regarding the most top dominant pathogens and provided additional information of resistance genes. Revised antibiotic treatment was followed by the patient's clinical recovery. Though sample preparation and the interpretation of final results still need to be improved further, metagenomic sequencing contributes to the accurate diagnosis of culture-negative infections and facilitates the rational antibiotic therapy.
Keywords: clinical diagnosis; metagenomic next-generation sequencing (mNGS); nanopore sequencing; severe pneumonia; whole genome sequencing.
Copyright © 2020 Wang, Li, Lin, Chen, Yang, Li, Zhang, Chen, Li, Du, Zhou, Li, Wang and Song.
Figures
Similar articles
-
Metagenomic identification of severe pneumonia pathogens in mechanically-ventilated patients: a feasibility and clinical validity study.Respir Res. 2019 Nov 27;20(1):265. doi: 10.1186/s12931-019-1218-4. Respir Res. 2019. PMID: 31775777 Free PMC article.
-
The application of nanopore targeted sequencing for pathogen diagnosis in bronchoalveolar lavage fluid of patients with pneumonia: a prospective multicenter study.Infect Dis (Lond). 2024 Feb;56(2):128-137. doi: 10.1080/23744235.2023.2276785. Epub 2023 Dec 18. Infect Dis (Lond). 2024. PMID: 37934028
-
Real-time analysis of nanopore-based metagenomic sequencing from infected orthopaedic devices.BMC Genomics. 2018 Sep 27;19(1):714. doi: 10.1186/s12864-018-5094-y. BMC Genomics. 2018. PMID: 30261842 Free PMC article.
-
Clinical Metagenomic Next-Generation Sequencing for Pathogen Detection.Annu Rev Pathol. 2019 Jan 24;14:319-338. doi: 10.1146/annurev-pathmechdis-012418-012751. Epub 2018 Oct 24. Annu Rev Pathol. 2019. PMID: 30355154 Free PMC article. Review.
-
Identification and characterization of CTX-M-15 producing Klebsiella pneumoniae clone ST101 in a Hungarian university teaching hospital.Acta Microbiol Immunol Hung. 2015 Sep;62(3):233-45. doi: 10.1556/030.62.2015.3.2. Acta Microbiol Immunol Hung. 2015. PMID: 26551567 Review.
Cited by
-
Clinical evaluation of metagenomic next-generation sequencing for detecting pathogens in bronchoalveolar lavage fluid collected from children with community-acquired pneumonia.Front Med (Lausanne). 2022 Jul 15;9:952636. doi: 10.3389/fmed.2022.952636. eCollection 2022. Front Med (Lausanne). 2022. PMID: 35911412 Free PMC article.
-
The Role of the Respiratory Microbiome in the Pathogenesis of Aspiration Pneumonia: Implications for Diagnosis and Potential Therapeutic Choices.Antibiotics (Basel). 2023 Jan 10;12(1):140. doi: 10.3390/antibiotics12010140. Antibiotics (Basel). 2023. PMID: 36671341 Free PMC article. Review.
-
Evaluation of 16S-Based Metagenomic NGS as Diagnostic Tool in Different Types of Culture-Negative Infections.Pathogens. 2024 Aug 30;13(9):743. doi: 10.3390/pathogens13090743. Pathogens. 2024. PMID: 39338934 Free PMC article.
-
Review of knockout technology approaches in bacterial drug resistance research.PeerJ. 2023 Aug 17;11:e15790. doi: 10.7717/peerj.15790. eCollection 2023. PeerJ. 2023. PMID: 37605748 Free PMC article. Review.
-
Application of Nanopore Sequencing in the Diagnosis and Treatment of Pulmonary Infections.Mol Diagn Ther. 2023 Nov;27(6):685-701. doi: 10.1007/s40291-023-00669-8. Epub 2023 Aug 11. Mol Diagn Ther. 2023. PMID: 37563539 Free PMC article. Review.
References
-
- CLSI (2019). Performance Standards for Antimicrobial Susceptibility Testing: Twenty-Nine Informational Supplement. M100, 29th Edn. Wayne, IL.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

