Deep learning-guided joint attenuation and scatter correction in multitracer neuroimaging studies
- PMID: 32436261
- PMCID: PMC7416024
- DOI: 10.1002/hbm.25039
Deep learning-guided joint attenuation and scatter correction in multitracer neuroimaging studies
Abstract
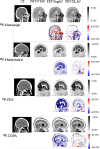
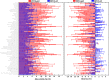
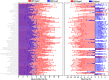
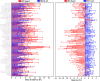
PET attenuation correction (AC) on systems lacking CT/transmission scanning, such as dedicated brain PET scanners and hybrid PET/MRI, is challenging. Direct AC in image-space, wherein PET images corrected for attenuation and scatter are synthesized from nonattenuation corrected PET (PET-nonAC) images in an end-to-end fashion using deep learning approaches (DLAC) is evaluated for various radiotracers used in molecular neuroimaging studies. One hundred eighty brain PET scans acquired using 18 F-FDG, 18 F-DOPA, 18 F-Flortaucipir (targeting tau pathology), and 18 F-Flutemetamol (targeting amyloid pathology) radiotracers (40 + 5, training/validation + external test, subjects for each radiotracer) were included. The PET data were reconstructed using CT-based AC (CTAC) to generate reference PET-CTAC and without AC to produce PET-nonAC images. A deep convolutional neural network was trained to generate PET attenuation corrected images (PET-DLAC) from PET-nonAC. The quantitative accuracy of this approach was investigated separately for each radiotracer considering the values obtained from PET-CTAC images as reference. A segmented AC map (PET-SegAC) containing soft-tissue and background air was also included in the evaluation. Quantitative analysis of PET images demonstrated superior performance of the DLAC approach compared to SegAC technique for all tracers. Despite the relatively low quantitative bias observed when using the DLAC approach, this approach appears vulnerable to outliers, resulting in noticeable local pseudo uptake and false cold regions. Direct AC in image-space using deep learning demonstrated quantitatively acceptable performance with less than 9% absolute SUV bias for the four different investigated neuroimaging radiotracers. However, this approach is vulnerable to outliers which result in large local quantitative bias.
Keywords: PET; attenuation correction; deep learning; neuroimaging tracers; quantification.
© 2020 The Authors. Human Brain Mapping published by Wiley Periodicals, Inc.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
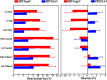

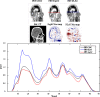
References
-
- Arabi, H. , Dowling, J. A. , Burgos, N. , Han, X. , Greer, P. B. , Koutsouvelis, N. , & Zaidi, H. (2018). Comparative study of algorithms for synthetic CT generation from MRI: Consequences for MRI‐guided radiation planning in the pelvic region. Medical Physics, 45, 5218–5233. - PubMed
-
- Arabi, H. , Koutsouvelis, N. , Rouzaud, M. , Miralbell, R. , & Zaidi, H. (2016). Atlas‐guided generation of pseudo‐CT images for MRI‐only and hybrid PET–MRI‐guided radiotherapy treatment planning. Physics in Medicine and Biology, 61, 6531–6552. - PubMed
-
- Arabi, H. , Rager, O. , Alem, A. , Varoquaux, A. , Becker, M. , & Zaidi, H. (2015). Clinical assessment of MR‐guided 3‐class and 4‐class attenuation correction in PET/MR. Molecular Imaging and Biology, 17, 264–276. - PubMed
-
- Arabi, H. , & Zaidi, H. (2016a). Magnetic resonance imaging‐guided attenuation correction in whole‐body PET/MRI using a sorted atlas approach. Medical Image Analysis, 31, 1–15. - PubMed
-
- Arabi, H. , & Zaidi, H. (2016b). One registration multi‐atlas‐based pseudo‐CT generation for attenuation correction in PET/MRI. European Journal of Nuclear Medicine and Molecular Imaging, 43, 2021–2035. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

