Pseudo-Improper-Dihedral Model for Intrinsically Disordered Proteins
- PMID: 32436706
- PMCID: PMC7588027
- DOI: 10.1021/acs.jctc.0c00338
Pseudo-Improper-Dihedral Model for Intrinsically Disordered Proteins
Abstract
We present a new coarse-grained Cα-based protein model with a nonradial multibody pseudo-improper-dihedral potential that is transferable, time-independent, and suitable for molecular dynamics. It captures the nature of backbone and side-chain interactions between amino acid residues by adapting a simple improper dihedral term for a one-bead-per-residue model. It is parameterized for intrinsically disordered proteins and applicable to simulations of such proteins and their assemblies on millisecond time scales.
Conflict of interest statement
The authors declare no competing financial interest.
Figures

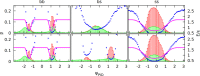
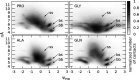
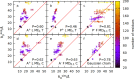
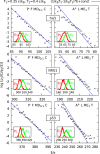
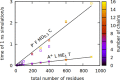
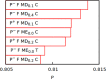
References
-
- Bixon M.; Lifson S. Potential functions and conformations in cycloalkanes. Tetrahedron 1967, 23, 769–784. 10.1016/0040-4020(67)85023-3. - DOI
-
- Lifson S.; Warshel A. Consistent force field for calculations of conformations, vibrational spectra, and enthalpies of cycloalkane and n-alkane molecules. J. Chem. Phys. 1968, 49, 5116–5129. 10.1063/1.1670007. - DOI
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
