A High-Throughput Screening System Based on Droplet Microfluidics for Glucose Oxidase Gene Libraries
- PMID: 32455903
- PMCID: PMC7287683
- DOI: 10.3390/molecules25102418
A High-Throughput Screening System Based on Droplet Microfluidics for Glucose Oxidase Gene Libraries
Abstract
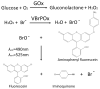
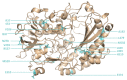
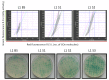
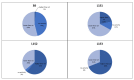
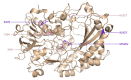
Glucose oxidase (GOx) is an important industrial enzyme that can be optimized for specific applications by mutagenesis and activity-based screening. To increase the efficiency of this approach, we have developed a new ultrahigh-throughput screening platform based on a microfluidic lab-on-chip device that allows the sorting of GOx mutants from a saturation mutagenesis library expressed on the surface of yeast cells. GOx activity was measured by monitoring the fluorescence of water microdroplets dispersed in perfluorinated oil. The signal was generated via a series of coupled enzyme reactions leading to the formation of fluorescein. Using this new method, we were able to enrich the yeast cell population by more than 35-fold for GOx mutants with higher than wild-type activity after two rounds of sorting, almost double the efficiency of our previously described flow cytometry platform. We identified and characterized novel GOx mutants, the most promising of which (M6) contained a combination of six point mutations that increased the catalytic constant kcat by 2.1-fold compared to wild-type GOx and by 1.4-fold compared to a parental GOx variant. The new microfluidic platform for GOx was therefore more sensitive than flow cytometry and supports comprehensive screens of gene libraries containing multiple mutations per gene.
Keywords: enzyme optimization; fluorescent label; protein engineering; sorting.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Development of GFP-based high-throughput screening system for directed evolution of glucose oxidase.J Biosci Bioeng. 2019 Jan;127(1):30-37. doi: 10.1016/j.jbiosc.2018.07.002. Epub 2018 Jul 20. J Biosci Bioeng. 2019. PMID: 30033354
-
Ultra-high-throughput screening method for the directed evolution of glucose oxidase.Chem Biol. 2014 Mar 20;21(3):414-21. doi: 10.1016/j.chembiol.2014.01.010. Epub 2014 Mar 6. Chem Biol. 2014. PMID: 24613019
-
Vanadium bromoperoxidase-coupled fluorescent assay for flow cytometry sorting of glucose oxidase gene libraries in double emulsions.Anal Bioanal Chem. 2012 Sep;404(5):1439-47. doi: 10.1007/s00216-012-6234-x. Epub 2012 Jul 19. Anal Bioanal Chem. 2012. PMID: 22811062
-
Ultrahigh-throughput FACS-based screening for directed enzyme evolution.Chembiochem. 2009 Nov 23;10(17):2704-15. doi: 10.1002/cbic.200900384. Chembiochem. 2009. PMID: 19780076 Review.
-
Speeding up enzyme discovery and engineering with ultrahigh-throughput methods.Curr Opin Struct Biol. 2018 Feb;48:149-156. doi: 10.1016/j.sbi.2017.12.010. Epub 2018 Feb 3. Curr Opin Struct Biol. 2018. PMID: 29413955 Review.
Cited by
-
Ultrahigh-Throughput Enzyme Engineering and Discovery in In Vitro Compartments.Chem Rev. 2023 May 10;123(9):5571-5611. doi: 10.1021/acs.chemrev.2c00910. Epub 2023 May 1. Chem Rev. 2023. PMID: 37126602 Free PMC article. Review.
-
Whole-cell screening of oxidative enzymes using genetically encoded sensors.Chem Sci. 2021 Oct 29;12(44):14766-14772. doi: 10.1039/d1sc02578c. eCollection 2021 Nov 17. Chem Sci. 2021. PMID: 34820092 Free PMC article.
-
Design, Screening, and Testing of Non-Rational Peptide Libraries with Antimicrobial Activity: In Silico and Experimental Approaches.Antibiotics (Basel). 2020 Nov 30;9(12):854. doi: 10.3390/antibiotics9120854. Antibiotics (Basel). 2020. PMID: 33265897 Free PMC article. Review.
-
Directed Evolution in Drops: Molecular Aspects and Applications.ACS Synth Biol. 2021 Nov 19;10(11):2772-2783. doi: 10.1021/acssynbio.1c00313. Epub 2021 Oct 22. ACS Synth Biol. 2021. PMID: 34677942 Free PMC article. Review.
-
Activity assays for flavoprotein oxidases: an overview.Appl Microbiol Biotechnol. 2025 May 8;109(1):115. doi: 10.1007/s00253-025-13494-2. Appl Microbiol Biotechnol. 2025. PMID: 40341429 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

