Rous Sarcoma Virus Genomic RNA Dimerization Capability In Vitro Is Not a Prerequisite for Viral Infectivity
- PMID: 32455905
- PMCID: PMC7291142
- DOI: 10.3390/v12050568
Rous Sarcoma Virus Genomic RNA Dimerization Capability In Vitro Is Not a Prerequisite for Viral Infectivity
Abstract
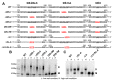
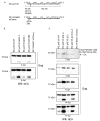
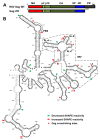
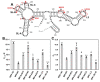
Retroviruses package their full-length, dimeric genomic RNA (gRNA) via specific interactions between the Gag polyprotein and a "Ψ" packaging signal located in the gRNA 5'-UTR. Rous sarcoma virus (RSV) gRNA has a contiguous, well-defined Ψ element, that directs the packaging of heterologous RNAs efficiently. The simplicity of RSV Ψ makes it an informative model to examine the mechanism of retroviral gRNA packaging, which is incompletely understood. Little is known about the structure of dimerization initiation sites or specific Gag interaction sites of RSV gRNA. Using selective 2'-hydroxyl acylation analyzed by primer extension (SHAPE), we probed the secondary structure of the entire RSV 5'-leader RNA for the first time. We identified a putative bipartite dimerization initiation signal (DIS), and mutation of both sites was required to significantly reduce dimerization in vitro. These mutations failed to reduce viral replication, suggesting that in vitro dimerization results do not strictly correlate with in vivo infectivity, possibly due to additional RNA interactions that maintain the dimers in cells. UV crosslinking-coupled SHAPE (XL-SHAPE) was next used to determine Gag-induced RNA conformational changes, revealing G218 as a critical Gag contact site. Overall, our results suggest that disruption of either of the DIS sequences does not reduce virus replication and reveal specific sites of Gag-RNA interactions.
Keywords: Gag polyprotein; RNA secondary structure; RNA structure-probing; RNA–protein interactions; Rous sarcoma virus (RSV); UV crosslinking-coupled SHAPE (XL-SHAPE); dimerization initiation signal (DIS); packaging signal (Ψ); retroviruses; selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE).
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Role of the Psi Packaging Signal and Dimerization Initiation Sequence in the Organization of Rous Sarcoma Virus Gag-gRNA Co-Condensates.Viruses. 2025 Jan 13;17(1):97. doi: 10.3390/v17010097. Viruses. 2025. PMID: 39861886 Free PMC article.
-
Visualizing Association of the Retroviral Gag Protein with Unspliced Viral RNA in the Nucleus.mBio. 2020 Apr 7;11(2):e00524-20. doi: 10.1128/mBio.00524-20. mBio. 2020. PMID: 32265329 Free PMC article.
-
Genetic evidence for a connection between Rous sarcoma virus gag nuclear trafficking and genomic RNA packaging.J Virol. 2009 Jul;83(13):6790-7. doi: 10.1128/JVI.00101-09. Epub 2009 Apr 15. J Virol. 2009. PMID: 19369339 Free PMC article.
-
On the Selective Packaging of Genomic RNA by HIV-1.Viruses. 2016 Sep 12;8(9):246. doi: 10.3390/v8090246. Viruses. 2016. PMID: 27626441 Free PMC article. Review.
-
Retroviral Gag protein-RNA interactions: Implications for specific genomic RNA packaging and virion assembly.Semin Cell Dev Biol. 2019 Feb;86:129-139. doi: 10.1016/j.semcdb.2018.03.015. Epub 2018 Apr 1. Semin Cell Dev Biol. 2019. PMID: 29580971 Free PMC article. Review.
Cited by
-
Visualizing Rous Sarcoma Virus Genomic RNA Dimerization in the Nucleus, Cytoplasm, and at the Plasma Membrane.Viruses. 2021 May 13;13(5):903. doi: 10.3390/v13050903. Viruses. 2021. PMID: 34068261 Free PMC article.
-
Two peas in a pod: retroviral RNA dimers organize Gag-RNA nanoclusters with novel biophysical properties.bioRxiv [Preprint]. 2025 Jun 11:2025.06.09.658653. doi: 10.1101/2025.06.09.658653. bioRxiv. 2025. Update in: Int J Mol Sci. 2025 Jun 13;26(12):5679. doi: 10.3390/ijms26125679. PMID: 40661432 Free PMC article. Updated. Preprint.
-
Correlation of SARS‑CoV‑2 to cancer: Carcinogenic or anticancer? (Review).Int J Oncol. 2022 Apr;60(4):42. doi: 10.3892/ijo.2022.5332. Epub 2022 Mar 2. Int J Oncol. 2022. PMID: 35234272 Free PMC article.
-
HIV-1 integrase binding to genomic RNA 5'-UTR induces local structural changes in vitro and in virio.Retrovirology. 2021 Nov 22;18(1):37. doi: 10.1186/s12977-021-00582-0. Retrovirology. 2021. PMID: 34809662 Free PMC article.
-
NMR Studies of Retroviral Genome Packaging.Viruses. 2020 Sep 30;12(10):1115. doi: 10.3390/v12101115. Viruses. 2020. PMID: 33008123 Free PMC article. Review.
References
-
- Brickell P.M. The p60c-src family of protein-tyrosine kinases: Structure, regulation, and function. Crit. Rev. Oncog. 1992;3:401–446. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

