The Epstein-Barr Virus Major Tegument Protein BNRF1 Is a Common Target of Cytotoxic CD4+ T Cells
- PMID: 32461311
- PMCID: PMC7375367
- DOI: 10.1128/JVI.00284-20
The Epstein-Barr Virus Major Tegument Protein BNRF1 Is a Common Target of Cytotoxic CD4+ T Cells
Abstract
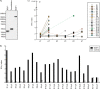
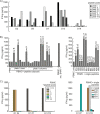
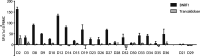
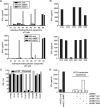
Cellular immunotherapy is a proven approach against Epstein-Barr virus (EBV)-driven lymphoproliferation in recipients of hematopoietic stem cells. Extending the applicability and improving the response rates of such therapy demands improving the knowledge base. We studied 23 healthy donors for specific CD4+ T cell responses against the viral tegument protein BNRF1 and found such T cells in all seropositive donors, establishing BNRF1 as an important immune target in EBV. We identified 18 novel immune epitopes from BNRF1, all of them generated by natural processing of the full-length protein from virus-transformed lymphoblastoid cell lines (LCL). BNRF1-specific CD4+ T cells were measured directly ex vivo by a cytokine-based method, thus providing a tool to study the interaction between immunity and infection in health and disease. T cells of the cytotoxic Th1 type inhibited the proliferation of autologous LCL as well as virus-driven transformation. We infer that they are important in limiting reactivations to subclinical levels during health and reducing virus propagation during disease. The information obtained from this work will feed into data sets that are indispensable in the design of patient-tailored immunotherapeutic approaches, thereby enabling the stride toward broader application of T cell therapy and improving clinical response rates.IMPORTANCE Epstein-Barr virus is carried by most humans and can cause life-threatening diseases. Virus-specific T cells have been used in different clinical settings with variable success rates. One way to improve immunotherapy is to better suit T cell generation protocols to viral targets available in different diseases. BNRF1 is present in viral particles and therefore likely available as a target for T cells in diseases with virus amplification. Here, we studied healthy Epstein-Barr virus (EBV) carriers for BNRF1 immunogenicity and report our results indicating BNRF1 to be a dominant target of the EBV-specific CD4+ T cell response. BNRF1-specific CD4+ T cells were found to be cytotoxic and capable of limiting EBV-driven B cell transformation in vitro The findings of this work contribute to forwarding our understanding of host-virus interactions during health and disease and are expected to find direct application in the generation of specific T cells for immunotherapy.
Keywords: Epstein-Barr virus; T cells; antigen; cytotoxic CD4+; virion structure.
Copyright © 2020 American Society for Microbiology.
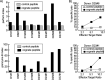
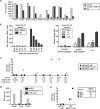
Figures

Similar articles
-
Combination of Epstein-Barr virus nuclear antigen 1, 3 and lytic antigen BZLF1 peptide pools allows fast and efficient stimulation of Epstein-Barr virus-specific T cells for adoptive immunotherapy.Cytotherapy. 2014 Jan;16(1):122-34. doi: 10.1016/j.jcyt.2013.07.008. Epub 2013 Oct 1. Cytotherapy. 2014. PMID: 24094498
-
In vitro cytokine production and growth inhibition of lymphoblastoid cell lines by CD4+ T cells from Epstein-Barr virus (EBV) seropositive donors.Clin Exp Immunol. 2001 Oct;126(1):101-10. doi: 10.1046/j.1365-2249.2001.01641.x. Clin Exp Immunol. 2001. PMID: 11678905 Free PMC article.
-
T cells specific for different latent and lytic viral proteins efficiently control Epstein-Barr virus-transformed B cells.Cytotherapy. 2015 Sep;17(9):1280-91. doi: 10.1016/j.jcyt.2015.06.003. Cytotherapy. 2015. PMID: 26276009
-
Immunodominance of lytic cycle antigens in Epstein-Barr virus-specific CD4+ T cell preparations for therapy.PLoS One. 2007 Jul 4;2(7):e583. doi: 10.1371/journal.pone.0000583. PLoS One. 2007. PMID: 17611619 Free PMC article. Review.
-
Immunodominant CD8 T cell response to Epstein-Barr virus.Biomed Pharmacother. 2001 Sep;55(7):373-80. doi: 10.1016/s0753-3322(01)00082-8. Biomed Pharmacother. 2001. PMID: 11669500 Review.
Cited by
-
Epstein-Barr virus BNRF1 destabilizes SMC5/6 cohesin complexes to evade its restriction of replication compartments.Cell Rep. 2022 Mar 8;38(10):110411. doi: 10.1016/j.celrep.2022.110411. Cell Rep. 2022. PMID: 35263599 Free PMC article.
-
Cytotoxic CD4+ T-cells specific for EBV capsid antigen BORF1 are maintained in long-term latently infected healthy donors.PLoS Pathog. 2021 Dec 9;17(12):e1010137. doi: 10.1371/journal.ppat.1010137. eCollection 2021 Dec. PLoS Pathog. 2021. PMID: 34882759 Free PMC article.
-
Functional Implications of Epstein-Barr Virus Lytic Genes in Carcinogenesis.Cancers (Basel). 2022 Nov 24;14(23):5780. doi: 10.3390/cancers14235780. Cancers (Basel). 2022. PMID: 36497262 Free PMC article. Review.
-
Evaluation of novel Epstein-Barr virus-derived antigen formulations for monitoring virus-specific T cells in pediatric patients with infectious mononucleosis.Virol J. 2024 Jun 14;21(1):139. doi: 10.1186/s12985-024-02411-0. Virol J. 2024. PMID: 38877590 Free PMC article.
-
Epstein-Barr virus lytic gene BNRF1 promotes B-cell lymphomagenesis via IFI27 upregulation.PLoS Pathog. 2024 Feb 1;20(2):e1011954. doi: 10.1371/journal.ppat.1011954. eCollection 2024 Feb. PLoS Pathog. 2024. PMID: 38300891 Free PMC article.
References
-
- IARC. 2012. A review of human carcinogens. Part B: biological agents, vol 100 B, p 475 International Agency for Research on Cancer, Lyon, France.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

