Association of microtubules and axonal RNA transferred from myelinating Schwann cells in rat sciatic nerve
- PMID: 32469980
- PMCID: PMC7259579
- DOI: 10.1371/journal.pone.0233651
Association of microtubules and axonal RNA transferred from myelinating Schwann cells in rat sciatic nerve
Abstract
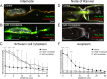
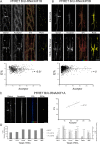
Transference of RNAs and ribosomes from Schwann cell-to-axon was demonstrated in normal and regenerating peripheral nerves. Previously, we have shown that RNAs transfer is dependent on F-actin cytoskeleton and Myosin Va. Here, we explored the contribution of microtubules to newly synthesized RNAs transport from Schwann cell nuclei up to nodal microvilli in sciatic nerves. Results using immunohistochemistry and quantitative confocal FRET analysis indicate that Schwann cell-derived RNAs co-localize with microtubules in Schwann cell cytoplasm. Additionally, transport of Schwann cell-derived RNAs is nocodazole and colchicine sensitive demonstrating its dependence on microtubule network integrity. Moreover, mRNAs codifying neuron-specific proteins are among Schwann cell newly synthesized RNAs population, and some of them are associated with KIF1B and KIF5B microtubules-based motors.
Conflict of interest statement
The authors declare that there is no conflict of interest regarding the publication of this article.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

