Cytoplasmic TDP43 Binds microRNAs: New Disease Targets in Amyotrophic Lateral Sclerosis
- PMID: 32477070
- PMCID: PMC7235295
- DOI: 10.3389/fncel.2020.00117
Cytoplasmic TDP43 Binds microRNAs: New Disease Targets in Amyotrophic Lateral Sclerosis
Abstract
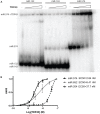
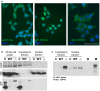
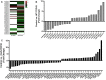
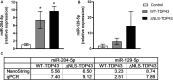
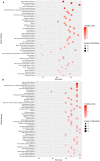
Amyotrophic lateral sclerosis (ALS) is a progressive, fatal, and incurable neurodegenerative disease. Recent studies suggest that dysregulation of gene expression by microRNAs (miRNAs) may play an important role in ALS pathogenesis. The reversible nature of this dysregulation makes miRNAs attractive pharmacological targets and a potential therapeutic avenue. Under physiological conditions, miRNA biogenesis, which begins in the nucleus and includes further maturation in the cytoplasm, involves trans-activation response element DNA/RNA-binding protein of 43 kDa (TDP43). However, TDP43 mutations or stress trigger TDP43 mislocalization and inclusion formation, a hallmark of most ALS cases, that may lead to aberrant protein/miRNA interactions in the cytoplasm. Herein, we demonstrated that TDP43 exhibits differential binding affinity for select miRNAs, which prompted us to profile miRNAs that preferentially bind cytoplasmic TDP43. Using cellular models expressing TDP43 variants and miRNA profiling analyses, we identified differential levels of 65 cytoplasmic TDP43-associated miRNAs. Of these, approximately 30% exhibited levels that differed by more than 3-fold in the cytoplasmic TDP43 models relative to our control model. The hits included both novel miRNAs and miRNAs previously associated with ALS that potentially regulate several predicted genes and pathways that may be important for pathogenesis. Accordingly, these findings highlight specific miRNAs that may shed light on relevant disease pathways and could represent potential biomarkers and reversible treatment targets for ALS.
Keywords: amyotrophic lateral sclerosis; cytoplasmic aggregates; microRNAs; profiling; trans-activation response element DNA/RNA-binding protein of 43 kDa (TDP43).
Copyright © 2020 Paez-Colasante, Figueroa-Romero, Rumora, Hur, Mendelson, Hayes, Backus, Taubman, Heinicke, Walter, Barmada, Sakowski and Feldman.
Figures
Similar articles
-
MicroRNAs: newcomers into the ALS picture.CNS Neurol Disord Drug Targets. 2015;14(2):194-207. doi: 10.2174/1871527314666150116125506. CNS Neurol Disord Drug Targets. 2015. PMID: 25613506 Review.
-
MicroRNA-183-5p regulates TAR DNA-binding protein 43 neurotoxicity via SQSTM1/p62 in amyotrophic lateral sclerosis.J Neurochem. 2023 Mar;164(5):643-657. doi: 10.1111/jnc.15744. Epub 2023 Jan 21. J Neurochem. 2023. PMID: 36527420
-
TDP43 nuclear export and neurodegeneration in models of amyotrophic lateral sclerosis and frontotemporal dementia.Sci Rep. 2018 Mar 15;8(1):4606. doi: 10.1038/s41598-018-22858-w. Sci Rep. 2018. PMID: 29545601 Free PMC article.
-
TDP43 and RNA instability in amyotrophic lateral sclerosis.Brain Res. 2018 Aug 15;1693(Pt A):67-74. doi: 10.1016/j.brainres.2018.01.015. Epub 2018 Jan 31. Brain Res. 2018. PMID: 29395044 Free PMC article. Review.
-
Interaction of RNA with a C-terminal fragment of the amyotrophic lateral sclerosis-associated TDP43 reduces cytotoxicity.Sci Rep. 2016 Jan 13;6:19230. doi: 10.1038/srep19230. Sci Rep. 2016. PMID: 26757674 Free PMC article.
Cited by
-
TDP-43 protein interactome informs about perturbed canonical pathways and may help develop personalized medicine approaches for patients with TDP-43 pathology.Drug Discov Today. 2023 Nov;28(11):103769. doi: 10.1016/j.drudis.2023.103769. Epub 2023 Sep 14. Drug Discov Today. 2023. PMID: 37714405 Free PMC article. Review.
-
Systematic review and meta-analysis of dysregulated microRNAs derived from liquid biopsies as biomarkers for amyotrophic lateral sclerosis.Noncoding RNA Res. 2024 Feb 6;9(2):523-535. doi: 10.1016/j.ncrna.2024.02.006. eCollection 2024 Jun. Noncoding RNA Res. 2024. PMID: 38511059 Free PMC article.
-
Dysregulation of MicroRNAs and PIWI-Interacting RNAs in a Caenorhabditis elegans Parkinson's Disease Model Overexpressing Human α-Synuclein and Influence of tdp-1.Front Neurosci. 2021 Mar 8;15:600462. doi: 10.3389/fnins.2021.600462. eCollection 2021. Front Neurosci. 2021. PMID: 33762903 Free PMC article.
-
Diagnostic Circulating miRNAs in Sporadic Amyotrophic Lateral Sclerosis.Front Med (Lausanne). 2022 May 6;9:861960. doi: 10.3389/fmed.2022.861960. eCollection 2022. Front Med (Lausanne). 2022. PMID: 35602517 Free PMC article.
-
Targeting TDP-43 Proteinopathy in hiPSC-Derived Mutated hNPCs with Mitoxantrone Drugs and miRNAs.Pharmaceutics. 2025 Mar 25;17(4):410. doi: 10.3390/pharmaceutics17040410. Pharmaceutics. 2025. PMID: 40284406 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

