Genetic Diversity and Evolutionary Analyses Reveal the Powdery Mildew Resistance Gene Pm21 Undergoing Diversifying Selection
- PMID: 32477413
- PMCID: PMC7241504
- DOI: 10.3389/fgene.2020.00489
Genetic Diversity and Evolutionary Analyses Reveal the Powdery Mildew Resistance Gene Pm21 Undergoing Diversifying Selection
Abstract
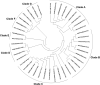
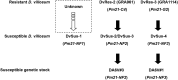
Wheat powdery mildew caused by Blumeria graminis f. sp. tritici (Bgt) is a devastating disease that threatens wheat production and yield worldwide. The powdery mildew resistance gene Pm21, originating from wheat wild relative Dasypyrum villosum, encodes a coiled-coil, nucleotide-binding site, leucine-rich repeat (CC-NBS-LRR) protein and confers broad-spectrum resistance to wheat powdery mildew. In the present study, we isolated 73 Pm21 alleles from different powdery mildew-resistant D. villosum accessions, among which, 38 alleles were non-redundant. Sequence analysis identified seven minor insertion-deletion (InDel) polymorphisms and 400 single nucleotide polymorphisms (SNPs) among the 38 non-redundant Pm21 alleles. The nucleotide diversity of the LRR domain was significantly higher than those of the CC and NB-ARC domains. Further evolutionary analysis indicated that the solvent-exposed LRR residues of Pm21 alleles had undergone diversifying selection (dN/dS = 3.19734). In addition, eight LRR motifs and four amino acid sites in the LRR domain were also experienced positive selection, indicating that these motifs and sites play critical roles in resistance specificity. The phylogenetic tree showed that 38 Pm21 alleles were divided into seven classes. Classes A (including original Pm21), B and C were the major classes, including 26 alleles (68.4%). We also identified three non-functional Pm21 alleles from four susceptible homozygous D. villosum lines (DvSus-1 to DvSus-4) and two susceptible wheat-D. villosum chromosome addition lines (DA6V#1 and DA6V#3). The genetic variations of non-functional Pm21 alleles involved point mutation, deletion and insertion, respectively. The results also showed that the non-functional Pm21 alleles in the two chromosome addition lines both came from the susceptible donors of D. villosum. This study gives a new insight into the evolutionary characteristics of Pm21 alleles and discusses how to sustainably utilize Pm21 in wheat production. This study also reveals the sequence variants and origins of non-functional Pm21 alleles in D. villosum populations.
Keywords: Dasypyrum villosum; Pm21 allele; diversifying selection; evolutionary analysis; genetic diversity; wheat powdery mildew resistance.
Copyright © 2020 He, Ji, Li, Tong, Feng, Wang, Han, Bie, Liu and Zhu.
Figures


Similar articles
-
Genetic, Physical and Comparative Mapping of the Powdery Mildew Resistance Gene Pm21 Originating from Dasypyrum villosum.Front Plant Sci. 2017 Nov 7;8:1914. doi: 10.3389/fpls.2017.01914. eCollection 2017. Front Plant Sci. 2017. PMID: 29163626 Free PMC article.
-
Screening and functional characterization of candidate resistance genes to powdery mildew from Dasypyrum villosum#4 in a wheat line Pm97033.Theor Appl Genet. 2020 Nov;133(11):3067-3083. doi: 10.1007/s00122-020-03655-4. Epub 2020 Jul 19. Theor Appl Genet. 2020. PMID: 32685983
-
Orthologous genes Pm12 and Pm21 from two wild relatives of wheat show evolutionary conservation but divergent powdery mildew resistance.Plant Commun. 2023 Mar 13;4(2):100472. doi: 10.1016/j.xplc.2022.100472. Epub 2022 Nov 9. Plant Commun. 2023. PMID: 36352792 Free PMC article.
-
Large-scale mutational analysis of wheat powdery mildew resistance gene Pm21.Front Plant Sci. 2022 Aug 9;13:988641. doi: 10.3389/fpls.2022.988641. eCollection 2022. Front Plant Sci. 2022. PMID: 36017260 Free PMC article.
-
Wheat powdery mildew resistance: from gene identification to immunity deployment.Front Plant Sci. 2023 Sep 18;14:1269498. doi: 10.3389/fpls.2023.1269498. eCollection 2023. Front Plant Sci. 2023. PMID: 37790783 Free PMC article. Review.
Cited by
-
Genetic mapping of powdery mildew resistance genes in wheat landrace Guizi 1 via genotyping by sequencing.Mol Biol Rep. 2022 Jun;49(6):4461-4468. doi: 10.1007/s11033-022-07287-3. Epub 2022 Mar 4. Mol Biol Rep. 2022. PMID: 35244868
-
Chromosomal mapping of a locus associated with adult-stage resistance to powdery mildew from Agropyron cristatum chromosome 6PL in wheat.Theor Appl Genet. 2022 Aug;135(8):2861-2873. doi: 10.1007/s00122-022-04155-3. Epub 2022 Jul 12. Theor Appl Genet. 2022. PMID: 35819492
-
Systematic Analysis of NB-ARC Gene Family in Rice and Functional Characterization of GNP12.Front Genet. 2022 Jun 16;13:887217. doi: 10.3389/fgene.2022.887217. eCollection 2022. Front Genet. 2022. PMID: 35783267 Free PMC article.
-
Physical mapping of a new powdery mildew resistance locus from Thinopyrum ponticum chromosome 4AgS.Front Plant Sci. 2023 Feb 23;14:1131205. doi: 10.3389/fpls.2023.1131205. eCollection 2023. Front Plant Sci. 2023. PMID: 36909389 Free PMC article.
-
Chromosome diversity in Dasypyrum villosum, an important genetic and trait resource for hexaploid wheat engineering.Ann Bot. 2023 Feb 7;131(1):185-198. doi: 10.1093/aob/mcac054. Ann Bot. 2023. PMID: 35451455 Free PMC article.
References
-
- Bie T., Zhao R., Jiang Z., Gao D., Zhang B., He H. (2015a). Efficient marker-assisted screening of structural changes involving Haynaldia villosa chromosome 6V using a double-distal-marker strategy. Mol. Breeding 35:34 10.1007/s11032-015-0211-y - DOI
-
- Bie T., Zhao R., Zhu S., Chen S., Cen B., Zhang B., et al. (2015b). Development and characterization of an efficient breeding-practical marker MBH1 simultaneously tagging Pm21 and PmV genes conferring resistance to wheat powdery mildew. Mol. Breeding 35:189 10.1007/s11032-015-0385-3 - DOI
-
- Cheng B., Ding Y., Gao X., Cao N., Xin Z., Zhang L. (2020). The diversity of powdery mildew resistance gene loci among wheat germplasm in Southwest China. Cereal Res. Commun. 48, 65–70. 10.1007/s42976-020-00015-2 - DOI
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

