Mitochondrial OXPHOS Biogenesis: Co-Regulation of Protein Synthesis, Import, and Assembly Pathways
- PMID: 32481479
- PMCID: PMC7312649
- DOI: 10.3390/ijms21113820
Mitochondrial OXPHOS Biogenesis: Co-Regulation of Protein Synthesis, Import, and Assembly Pathways
Abstract
The assembly of mitochondrial oxidative phosphorylation (OXPHOS) complexes is an intricate process, which-given their dual-genetic control-requires tight co-regulation of two evolutionarily distinct gene expression machineries. Moreover, fine-tuning protein synthesis to the nascent assembly of OXPHOS complexes requires regulatory mechanisms such as translational plasticity and translational activators that can coordinate mitochondrial translation with the import of nuclear-encoded mitochondrial proteins. The intricacy of OXPHOS complex biogenesis is further evidenced by the requirement of many tightly orchestrated steps and ancillary factors. Early-stage ancillary chaperones have essential roles in coordinating OXPHOS assembly, whilst late-stage assembly factors-also known as the LYRM (leucine-tyrosine-arginine motif) proteins-together with the mitochondrial acyl carrier protein (ACP)-regulate the incorporation and activation of late-incorporating OXPHOS subunits and/or co-factors. In this review, we describe recent discoveries providing insights into the mechanisms required for optimal OXPHOS biogenesis, including the coordination of mitochondrial gene expression with the availability of nuclear-encoded factors entering via mitochondrial protein import systems.
Keywords: LYRM proteins; OXPHOS assembly factors; OXPHOS biogenesis; mitochondrial ACP; mitochondrial gene expression; mitochondrial import.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
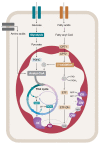

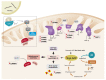
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

