PICRUSt2 for prediction of metagenome functions
- PMID: 32483366
- PMCID: PMC7365738
- DOI: 10.1038/s41587-020-0548-6
PICRUSt2 for prediction of metagenome functions
Figures
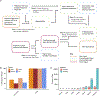

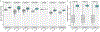
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

