Current approaches for automated model building into cryo-EM maps using Buccaneer with CCP-EM
- PMID: 32496215
- PMCID: PMC7271950
- DOI: 10.1107/S2059798320005513
Current approaches for automated model building into cryo-EM maps using Buccaneer with CCP-EM
Abstract
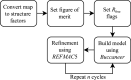
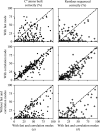
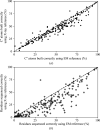
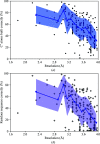
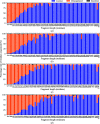
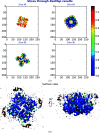
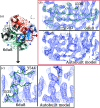
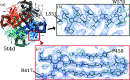
This work focuses on the use of the existing protein-model-building software Buccaneer to provide structural interpretation of electron cryo-microscopy (cryo-EM) maps. Originally developed for application to X-ray crystallography, the necessary steps to optimise the usage of Buccaneer with cryo-EM maps are shown. This approach has been applied to the data sets of 208 cryo-EM maps with resolutions of better than 4 Å. The results obtained also show an evident improvement in the sequencing step when the initial reference map and model used for crystallographic cases are replaced by a cryo-EM reference. All other necessary changes to settings in Buccaneer are implemented in the model-building pipeline from within the CCP-EM interface (as of version 1.4.0).
Keywords: Buccaneer; CCP-EM; Collaborative Computational Project for Electron cryo-Microscopy; cryo-EM; model building.
open access.
Figures

Similar articles
-
The accuracy of protein models automatically built into cryo-EM maps with ARP/wARP.Acta Crystallogr D Struct Biol. 2021 Feb 1;77(Pt 2):142-150. doi: 10.1107/S2059798320016332. Epub 2021 Jan 26. Acta Crystallogr D Struct Biol. 2021. PMID: 33559604 Free PMC article.
-
Current approaches for the fitting and refinement of atomic models into cryo-EM maps using CCP-EM.Acta Crystallogr D Struct Biol. 2018 Jun 1;74(Pt 6):492-505. doi: 10.1107/S2059798318007313. Epub 2018 May 30. Acta Crystallogr D Struct Biol. 2018. PMID: 29872001 Free PMC article. Review.
-
Redeployment of automated MrBUMP search-model identification for map fitting in cryo-EM.Acta Crystallogr D Struct Biol. 2021 Nov 1;77(Pt 11):1378-1385. doi: 10.1107/S2059798321009165. Epub 2021 Oct 20. Acta Crystallogr D Struct Biol. 2021. PMID: 34726166 Free PMC article.
-
New tools for the analysis and validation of cryo-EM maps and atomic models.Acta Crystallogr D Struct Biol. 2018 Sep 1;74(Pt 9):814-840. doi: 10.1107/S2059798318009324. Epub 2018 Sep 3. Acta Crystallogr D Struct Biol. 2018. PMID: 30198894 Free PMC article.
-
Refinement of Atomic Structures Against cryo-EM Maps.Methods Enzymol. 2016;579:277-305. doi: 10.1016/bs.mie.2016.05.033. Epub 2016 Jun 24. Methods Enzymol. 2016. PMID: 27572731 Review.
Cited by
-
ModelCraft: an advanced automated model-building pipeline using Buccaneer.Acta Crystallogr D Struct Biol. 2022 Sep 1;78(Pt 9):1090-1098. doi: 10.1107/S2059798322007732. Epub 2022 Aug 25. Acta Crystallogr D Struct Biol. 2022. PMID: 36048149 Free PMC article.
-
Cryo-EM structures of pentameric autoinducer-2 exporter from Escherichia coli reveal its transport mechanism.EMBO J. 2022 Sep 15;41(18):e109990. doi: 10.15252/embj.2021109990. Epub 2022 Jun 14. EMBO J. 2022. PMID: 35698912 Free PMC article.
-
findMySequence: a neural-network-based approach for identification of unknown proteins in X-ray crystallography and cryo-EM.IUCrJ. 2021 Dec 1;9(Pt 1):86-97. doi: 10.1107/S2052252521011088. eCollection 2022 Jan 1. IUCrJ. 2021. PMID: 35059213 Free PMC article.
-
DoubleHelix: nucleic acid sequence identification, assignment and validation tool for cryo-EM and crystal structure models.Nucleic Acids Res. 2023 Aug 25;51(15):8255-8269. doi: 10.1093/nar/gkad553. Nucleic Acids Res. 2023. PMID: 37395405 Free PMC article.
-
Large language models generate functional protein sequences across diverse families.Nat Biotechnol. 2023 Aug;41(8):1099-1106. doi: 10.1038/s41587-022-01618-2. Epub 2023 Jan 26. Nat Biotechnol. 2023. PMID: 36702895 Free PMC article.

