A Proximity Mapping Journey into the Biology of the Mammalian Centrosome/Cilium Complex
- PMID: 32503249
- PMCID: PMC7348975
- DOI: 10.3390/cells9061390
A Proximity Mapping Journey into the Biology of the Mammalian Centrosome/Cilium Complex
Abstract
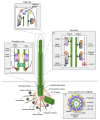
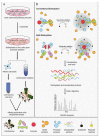
The mammalian centrosome/cilium complex is composed of the centrosome, the primary cilium and the centriolar satellites, which together regulate cell polarity, signaling, proliferation and motility in cells and thereby development and homeostasis in organisms. Accordingly, deregulation of its structure and functions is implicated in various human diseases including cancer, developmental disorders and neurodegenerative diseases. To better understand these disease connections, the molecular underpinnings of the assembly, maintenance and dynamic adaptations of the centrosome/cilium complex need to be uncovered with exquisite detail. Application of proximity-based labeling methods to the centrosome/cilium complex generated spatial and temporal interaction maps for its components and provided key insights into these questions. In this review, we first describe the structure and cell cycle-linked regulation of the centrosome/cilium complex. Next, we explain the inherent biochemical and temporal limitations in probing the structure and function of the centrosome/cilium complex and describe how proximity-based labeling approaches have addressed them. Finally, we explore current insights into the knowledge we gained from the proximity mapping studies as it pertains to centrosome and cilium biogenesis and systematic characterization of the centrosome, cilium and centriolar satellite interactomes.
Keywords: APEX; BioID; TurboID; centriolar satellites; centrosome; cilia; ciliopathies; microtubules; proximity-labeling.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

