A substitution in cGMP-dependent protein kinase 1 associated with aortic disease induces an active conformation in the absence of cGMP
- PMID: 32506052
- PMCID: PMC7383375
- DOI: 10.1074/jbc.RA119.010984
A substitution in cGMP-dependent protein kinase 1 associated with aortic disease induces an active conformation in the absence of cGMP
Abstract
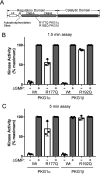
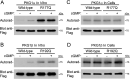
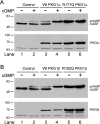
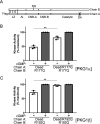
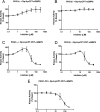
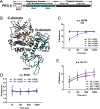
Type 1 cGMP-dependent protein kinases (PKGs) play important roles in human cardiovascular physiology, regulating vascular tone and smooth-muscle cell phenotype. A mutation in the human PRKG1 gene encoding cGMP-dependent protein kinase 1 (PKG1) leads to thoracic aortic aneurysms and dissections. The mutation causes an arginine-to-glutamine (RQ) substitution within the first cGMP-binding pocket in PKG1. This substitution disrupts cGMP binding to the pocket, but it also unexpectedly causes PKG1 to have high activity in the absence of cGMP via an unknown mechanism. Here, we identified the molecular mechanism whereby the RQ mutation increases basal kinase activity in the human PKG1α and PKG1β isoforms. Although we found that the RQ substitution (R177Q in PKG1α and R192Q in PKG1β) increases PKG1α and PKG1β autophosphorylation in vitro, we did not detect increased autophosphorylation of the PKG1α or PKG1β RQ variant isolated from transiently transfected 293T cells, indicating that increased basal activity of the RQ variants in cells was not driven by PKG1 autophosphorylation. Replacement of Arg-177 in PKG1α with alanine or methionine also increased basal activity. PKG1 exists as a parallel homodimer linked by an N-terminal leucine zipper, and we show that the WT chain in WT-RQ heterodimers partly reduces basal activity of the RQ chain. Using hydrogen/deuterium-exchange MS, we found that the RQ substitution causes PKG1β to adopt an active conformation in the absence of cGMP, similar to that of cGMP-bound WT enzyme. We conclude that the RQ substitution in PKG1 increases its basal activity by disrupting the formation of an inactive conformation.
Keywords: autophosphorylation; cardiovascular system; cyclic GMP (cGMP); enzyme structure; hydrogen exchange mass spectrometry; hydrogen/deuterium exchange; kinase signaling; mutagenesis; mutant; protein kinase G (PKG); thoracic aortic aneurysm and dissection (TAAD).
© 2020 Chan et al.
Conflict of interest statement
Conflict of interest—The authors declare no conflicts of interest with the content of this article.
Figures


References
-
- Friebe A., Sandner P., and Schmidtko A. (2017) Meeting report of the 8th International Conference on cGMP “cGMP: generators, effectors, and therapeutic implications” at Bamberg, Germany, from June 23 to 25, 2017. Naunyn. Schmiedebergs Arch. Pharmacol. 390, 1177–1188 10.1007/s00210-017-1429-5 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

