Identification of potential candidate genes and pathways in atrioventricular nodal reentry tachycardia by whole-exome sequencing
- PMID: 32508047
- PMCID: PMC7240861
- DOI: 10.1002/ctm2.25
Identification of potential candidate genes and pathways in atrioventricular nodal reentry tachycardia by whole-exome sequencing
Erratum in
-
ERRATUM.Clin Transl Med. 2021 Feb;11(2):e274. doi: 10.1002/ctm2.274. Clin Transl Med. 2021. PMID: 33635007 Free PMC article. No abstract available.
Abstract
Background: Atrioventricular nodal reentry tachycardia (AVNRT) is the most common manifestation of paroxysmal supraventricular tachycardia (PSVT). Increasing data have indicated familial clustering and participation of genetic factors in AVNRT, and no pathogenic genes related to AVNRT have been reported.
Methods: Whole-exome sequencing (WES) was performed in 82 patients with AVNRT and 100 controls. Reference genes, genome-wide association analysis, gene-based collapsing, and pathway enrichment analysis were performed. A protein-protein interaction (PPI) network was then established; WES database in the UK Biobank and one only genetic study of AVNRT in Denmark were used for external data validation.
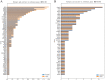
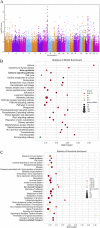
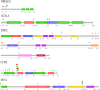
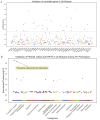
Results: Among 95 reference genes, 126 rare variants in 48 genes were identified in the cases (minor allele frequency < 0.001). Gene-based collapsing analysis and pathway enrichment analysis revealed six functional pathways related to AVNRT as with neuronal system/neurotransmitter release cycles and ion channel/cardiac conduction among the top 30 enriched pathways, and then 36 candidate pathogenic genes were selected. By combining with PPI analysis, 10 candidate genes were identified, including RYR2, NOS1, SCN1A, CFTR, EPHB4, ROBO1, PRKAG2, MMP2, ASPH, and ABCC8. From the UK Biobank database, 18 genes from candidate genes including SCN1A, PRKAG2, NOS1, and CFTR had rare variants in arrhythmias, and the rare variants in PIK3CB, GAD2, and HIP1R were in patients with PSVT. Moreover, one rare variant of RYR2 (c.4652A > G, p.Asn1551Ser) in our study was also detected in the Danish study. Considering the gene functional roles and external data validation, the most likely candidate genes were SCN1A, PRKAG2, RYR2, CFTR, NOS1, PIK3CB, GAD2, and HIP1R.
Conclusion: The preliminary results first revealed potential candidate genes such as SCN1A, PRKAG2, RYR2, CFTR, NOS1, PIK3CB, GAD2, and HIP1R, and the pathways mediated by these genes, including neuronal system/neurotransmitter release cycles or ion channels/cardiac conduction, might be involved in AVNRT.
Keywords: atrioventricular nodal reentry tachycardia, whole-exome sequencing; gene-based collapsing analysis, neurotransmitter release cycles pathway, ion channels-related pathway, ion channel genes.
© 2020 The Authors. Clinical and Translational Medicine published by John Wiley & Sons Australia, Ltd on behalf of Shanghai Institute of Clinical Bioinformatics.
Conflict of interest statement
The authors report no conflicts of interest. The authors alone are responsible for the content and writing of this article. The manuscript has been approved by the responsible authorities of the institutions where the work was conducted, and all authors have read the manuscript and approved its submission to your journal.
Figures


References
-
- Akhtar M, Jazayeri MR, Sra J, Blanck Z, Deshpande S, Dhala A. Atrioventricular nodal reentry: clinical, electrophysiological, and therapeutic considerations. Circulation. 1993;88:282‐295. - PubMed
-
- McGuire MA, Janse MJ. New insights on anatomical location of components of the reentrant circuit and ablation therapy for atrioventricular junctional reentrant tachycardia. Curr Opin Cardiol. 1995;10:3‐8. - PubMed
-
- Wu J, Zipes DP. Mechanisms underlying atrioventricular nodal conduction and the reentrant circuit of atrioventricular nodal reentrant tachycardia using optical mapping. J Cardiovasc Electrophysiol. 2002;13:831‐834. - PubMed
-
- Lu CW, Wu MH, Chu SH. Paroxysmal supraventricular tachycardia in identical twins with the same left lateral accessory pathways and innocent dual atrioventricular pathways. Pacing Clin Electrophysiol. 2000;23:1564‐1566. - PubMed
-
- Hayes JJ, Sharma PP, Smith PN, Vidaillet HJ. Familial atrioventricular nodal reentry tachycardia. Pacing Clin Electrophysiol. 2004;27:73‐76. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
