Measurements of Physiological Stress Responses in C. Elegans
- PMID: 32510480
- PMCID: PMC7840273
- DOI: 10.3791/61001
Measurements of Physiological Stress Responses in C. Elegans
Abstract
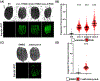
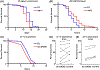
Organisms are often exposed to fluctuating environments and changes in intracellular homeostasis, which can have detrimental effects on their proteome and physiology. Thus, organisms have evolved targeted and specific stress responses dedicated to repair damage and maintain homeostasis. These mechanisms include the unfolded protein response of the endoplasmic reticulum (UPRER), the unfolded protein response of the mitochondria (UPRMT), the heat shock response (HSR), and the oxidative stress response (OxSR). The protocols presented here describe methods to detect and characterize the activation of these pathways and their physiological consequences in the nematode, C. elegans. First, the use of pathway-specific fluorescent transcriptional reporters is described for rapid cellular characterization, drug screening, or large-scale genetic screening (e.g., RNAi or mutant libraries). In addition, complementary, robust physiological assays are described, which can be used to directly assess sensitivity of animals to specific stressors, serving as functional validation of the transcriptional reporters. Together, these methods allow for rapid characterization of the cellular and physiological effects of internal and external proteotoxic perturbations.
Figures