A single Proteus mirabilis lineage from human and animal sources: a hidden reservoir of OXA-23 or OXA-58 carbapenemases in Enterobacterales
- PMID: 32514057
- PMCID: PMC7280188
- DOI: 10.1038/s41598-020-66161-z
A single Proteus mirabilis lineage from human and animal sources: a hidden reservoir of OXA-23 or OXA-58 carbapenemases in Enterobacterales
Abstract
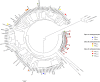
In Enterobacterales, the most common carbapenemases are Ambler's class A (KPC-like), class B (NDM-, VIM- or IMP-like) or class D (OXA-48-like) enzymes. This study describes the characterization of twenty-four OXA-23 or OXA-58 producing-Proteus mirabilis isolates recovered from human and veterinary samples from France and Belgium. Twenty-two P. mirabilis isolates producing either OXA-23 (n = 21) or OXA-58 (n = 1), collected between 2013 and 2018, as well as 2 reference strains isolated in 1996 and 2015 were fully sequenced. Phylogenetic analysis revealed that 22 of the 24 isolates, including the isolate from 1996, belonged to a single lineage that has disseminated in humans and animals over a long period of time. The blaOXA-23 gene was located on the chromosome and was part of a composite transposon, Tn6703, bracketed by two copies of IS15∆II. Sequencing using Pacbio long read technology of OXA-23-producing P. mirabilis VAC allowed the assembly of a 55.5-kb structure encompassing the blaOXA-23 gene in that isolate. By contrast to the blaOXA-23 genes, the blaOXA-58 gene of P. mirabilis CNR20130297 was identified on a 6-kb plasmid. The acquisition of the blaOXA-58 gene on this plasmid involved XerC-XerD recombinases. Our results suggest that a major clone of OXA-23-producing P. mirabilis is circulating in France and Belgium since 1996.
Conflict of interest statement
The authors declare no competing interests.
Figures


References
-
- Frontiers | Genetics of Acquired Antibiotic Resistance Genes in Proteus spp. | Microbiology, https://www.frontiersin.org/articles/10.3389/fmicb.2020.00256/full. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

