The DNA methylation landscape of hematological malignancies: an update
- PMID: 32526054
- PMCID: PMC7400809
- DOI: 10.1002/1878-0261.12744
The DNA methylation landscape of hematological malignancies: an update
Abstract
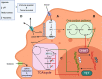
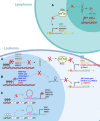
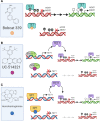
The rapid advances in high-throughput sequencing technologies have made it more evident that epigenetic modifications orchestrate a plethora of complex biological processes. During the last decade, we have gained significant knowledge about a wide range of epigenetic changes that crucially contribute to some of the most aggressive forms of leukemia, lymphoma, and myelodysplastic syndromes. DNA methylation is a key epigenetic player in the abnormal initiation, development, and progression of these malignancies, often acting in synergy with other epigenetic alterations. It also contributes to the acquisition of drug resistance. In this review, we summarize the role of DNA methylation in hematological malignancies described in the current literature. We discuss in detail the dual role of DNA methylation in normal and aberrant hematopoiesis, as well as the involvement of this type of epigenetic change in other aspects of the disease. Finally, we present a comprehensive overview of the main clinical implications, including a discussion of the therapeutic strategies that regulate or reverse aberrant DNA methylation patterns in hematological malignancies, including their combination with (chemo)immunotherapy.
Keywords: cancer; epigenetics; hematology; leukemia; lymphoma; methylation.
© 2020 The Authors. Published by FEBS Press and John Wiley & Sons Ltd.
Conflict of interest statement
ME is a consultant for Ferrer International and Quimatryx. The other authors declare that they have no conflict of interest.
Figures

References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

