A brainwide atlas of synapses across the mouse life span
- PMID: 32527927
- PMCID: PMC7115813
- DOI: 10.1126/science.aba3163
A brainwide atlas of synapses across the mouse life span
Abstract
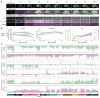
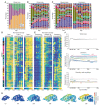
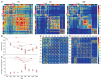
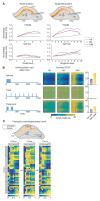
Synapses connect neurons together to form the circuits of the brain, and their molecular composition controls innate and learned behavior. We analyzed the molecular and morphological diversity of 5 billion excitatory synapses at single-synapse resolution across the mouse brain from birth to old age. A continuum of changes alters synapse composition in all brain regions across the life span. Expansion in synapse diversity produces differentiation of brain regions until early adulthood, and compositional changes cause dedifferentiation in old age. The spatiotemporal synaptome architecture of the brain potentially accounts for life-span transitions in intellectual ability, memory, and susceptibility to behavioral disorders.
Copyright © 2020 The Authors, some rights reserved; exclusive licensee American Association for the Advancement of Science. No claim to original U.S. Government Works.
Conflict of interest statement
Figures
Comment in
-
A synapse census for the ages.Science. 2020 Jul 17;369(6501):253-254. doi: 10.1126/science.abc9555. Science. 2020. PMID: 32675362 Free PMC article. No abstract available.
References
-
- Grant SG, et al. Impaired long-term potentiation, spatial learning, and hippocampal development in fyn mutant mice. Science. 1992;258:1903–1910. - PubMed
-
- Migaud M, et al. Enhanced long-term potentiation and impaired learning in mice with mutant postsynaptic density-95 protein. Nature. 1998;396:433–439. - PubMed
-
- Silva AJ, Paylor R, Wehner JM, Tonegawa S. Impaired spatial learning in alpha-calcium-calmodulin kinase II mutant mice. Science. 1992;257:206–211. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

